Sometimes, the most valuable thing you need on your journey to a healthy lifestyle is some guidance and motivation. Luckily, some user-friendly apps and gadgets can help you achieve your fitness goals in 2024.
The following devices are some of the newer products health enthusiasts are raving about this year. They are designed to track your progress, help you recover, keep you up to date on trends, and inform you about proper healthy habits. How many of these tools are you going to check out or add to your gym bag?
1. Lumen Metabolism Tracker


- Image Credit: Metaflow LTD.
The Lumen metabolism tracker allows users to blow into a sensor, which tracks the carbon dioxide concentration in their breath. This indicates whether their body is burning fat or carbohydrates. From there, it breaks down a daily nutritional plan to give you the optimal time to eat or fast. It can tell you if you are fasting too long and no longer burning fat or if you are comfortably shedding weight. If you have a goal of slimming down this year, this ground-breaking technology could be your answer.
2. Apollo Neuro Stress Relief Band


- Image Credit: Apollo Neuroscience, Inc.
The Apollo Neuro bracelet wraps around your wrist. It uses scientifically proven touch therapy to send tiny vibrations through your body. The goal is to calm your nervous system and improve your body’s reaction to stress triggers. Users have reported better quality of sleep, heightened focus, and lower levels of anxiety.
The device only needs to be worn when your body needs it. When you need to relax and unwind, this device is ideal for naturally training your body to deal with stress.
3. Fitbit Aria Air Scale


- Image Credit: Google LLC.
This smart scale syncs with your smartphone and tracks body weight and BMI while analyzing the data. It works with any Fitbit smartwatch and helps users gather more comprehensive data and trends about their health, workout routines, lifestyle, and body weight.
The scale can connect to multiple users to create a support system for people taking charge of their health. For as little as $40, this gadget is a must-have for fitness enthusiasts.
4. Molekule Air Purifier


- Image Credit: Molekule.
Whether you suffer from allergies or want to breathe the freshest air possible, this home air purifier is a life changer. The Molecule Air Purifier can easily and automatically clean the air in a room as big as 600 square feet.
The device comes with two separate filters. The first filter traps bigger particles like dust and pet hair, while the second breaks down pollutants at a molecular level. Bacteria, mold, viruses, allergens, and other contaminants don’t stand a chance of breaking through the proprietary light-activated catalyst technology this purifier boasts.
The device can be controlled by an app, sits quietly in the corner, and provides endless amounts of healthy air for you and your family.
5. MUSE S: The Brain Sensing Headband


- Image Credit: Muse.
Studies have proven that regular meditation can reduce stress, improve sleep quality, fight addiction, and lower blood pressure. The MUSE S is determined to make your meditation sessions that much better by tracking and analyzing your body’s measurements.
Worn across the user’s forehead, the MUSE S measures heart rate, breathing, subtle body movements, and brain waves. The MUSE app provides biofeedback in real-time. Users can also use the device to track sleep habits, assist in guided meditation, and perform breathing exercises.
6. Noom Weight Loss App
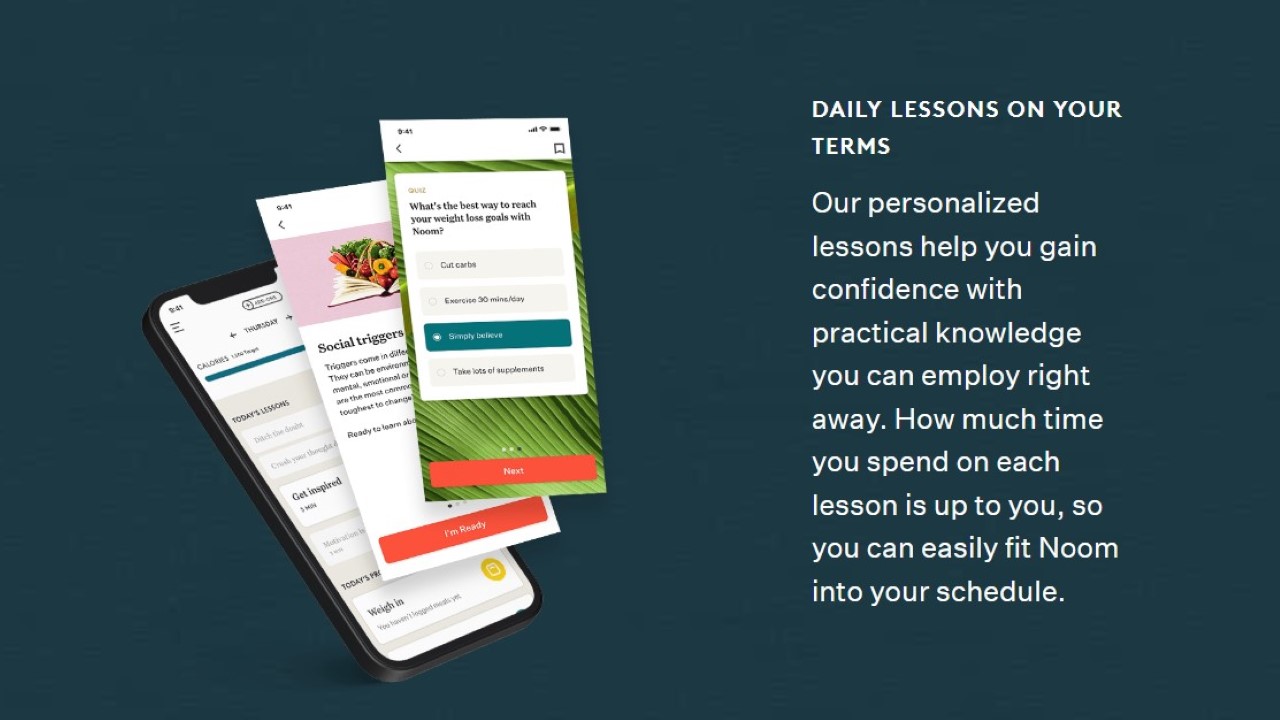

- Image Credit: Noom, Inc.
I’m sure some of you have seen the commercials for the weight loss app Noom. The brand’s approach to weight loss includes science and personalized programs to lose weight permanently. They believe in teaching their clients how to cope with their relationship with food, be conscious of their habits, and instill healthy lifestyle knowledge.
Daily lessons promote the company’s core beliefs. They want you to lose weight and understand why you are losing it. Food trackers and exercise logs are included to keep you consistent and on track to a healthier body.
A four-month subscription costs only $149, making it a fantastic resource for users looking to shed some pounds before summer.
7. Oral-B iO Series 9 Toothbrush


- Image Credit: Procter & Gamble.
It seems like every gadget we use daily is getting upgraded to a smarter version. This smart technology automatic toothbrush is designed to keep our teeth clean, kill bad breath, and brighten our smiles. The toothbrush pairs with the mobile app and assures the user that it hits 100% of their mouth with the proper pressure and length of time. The device will also inform you when to replace the brush head for optimal oral care.
8. Dr. Relief Acupressure Mat


- Image Credit: Dr. Relief.
I have personally never tried acupuncture, but many fitness experts swear by the results of this ancient Chinese medicine. Studies have shown that the practice of acupuncture can improve sleep, erase migraines, improve mental health, and temper chronic pain. Still, for some, the thought of needles in our bodies is beyond scary.
That is where this Dr. Relief mat comes in. It is thought to mimic the results without using those terrifying needles. The mat has a comfortable headrest that allows you to lie down for a full-body, acupuncture-like experience.
9. TheraGun Percussion Massager


- Image Credit: Therabody, Inc.
If you ask any personal trainer or fitness expert, they will tell you that recovery is just as important as the actual workout. Tired muscles need time to recover and grow before training again, and failure to do so can risk serious injury. Massages can be the ideal recovery tool for a sore body but can be expensive. The TheraGun percussion massager lets you get quality massages at home quickly and easily.
The machine provides various speeds and pressures and effectively works out knots and target spots. Its compact design makes it portable, so you can use it at home, in the office, or on vacation.
10. Oura Ring


- Image Credit: Ōura Health Oy.
The fashionable Oura Ring has built-in sensors to track and collect data 24 hours a day. It is quickly becoming one of the more advanced fitness trackers on the market. The third-generation Oura can successfully track sleep patterns, heart rate, body temperature, blood oxygen level, steps, distance traveled, calories burned, and downtime. The ring can also alert you if you are getting sick, experiencing high levels of stress, or need more sleep.
You might think a resource like this would cost a fortune. Nope. The ring has a price tag of $299, making it a great option for fitness fanatics or people looking to better understand their bodies.
11. QardioArm Wireless Smart Blood Pressure Monitor


- Image Credit: Qardio, Inc.
This QardioArm monitor takes the difficulty out of monitoring your blood pressure. The device wraps around your upper arm and instantly connects with your smartphone, making it super simple to send analyzed data to your medical provider.
The QardioArm is designed to measure systolic and diastolic blood pressure levels and irregular heartbeat. You can set reminders, geo-tracking, and a relaxation mode. It is compact and portable with a rechargeable battery, making it one of the most convenient blood pressure monitors on the market.
12. Fitbit Sense 2 Fitness & Health Tracker


- Image Credit: Google LLC.
Fitbit has continued to make high-quality fitness trackers, and the newest Sense 2 is no different. Not only is the futuristic case stylish and cool, but the technology has grown to help us store our fitness habits even better.
The watch is capable of tracking many bodily functions. It monitors heart rate, calories burned, steps, distance, elevation gain, and health trends. You can set the watch to different workout modes, rate your quality of sleep, and alert you to irregular heartbeats that could be a cause of an underlying health factor. All in all, this gadget is an amazing tool to have if you want to be informed of your body’s actions at all times.
14. Apple Fitness +
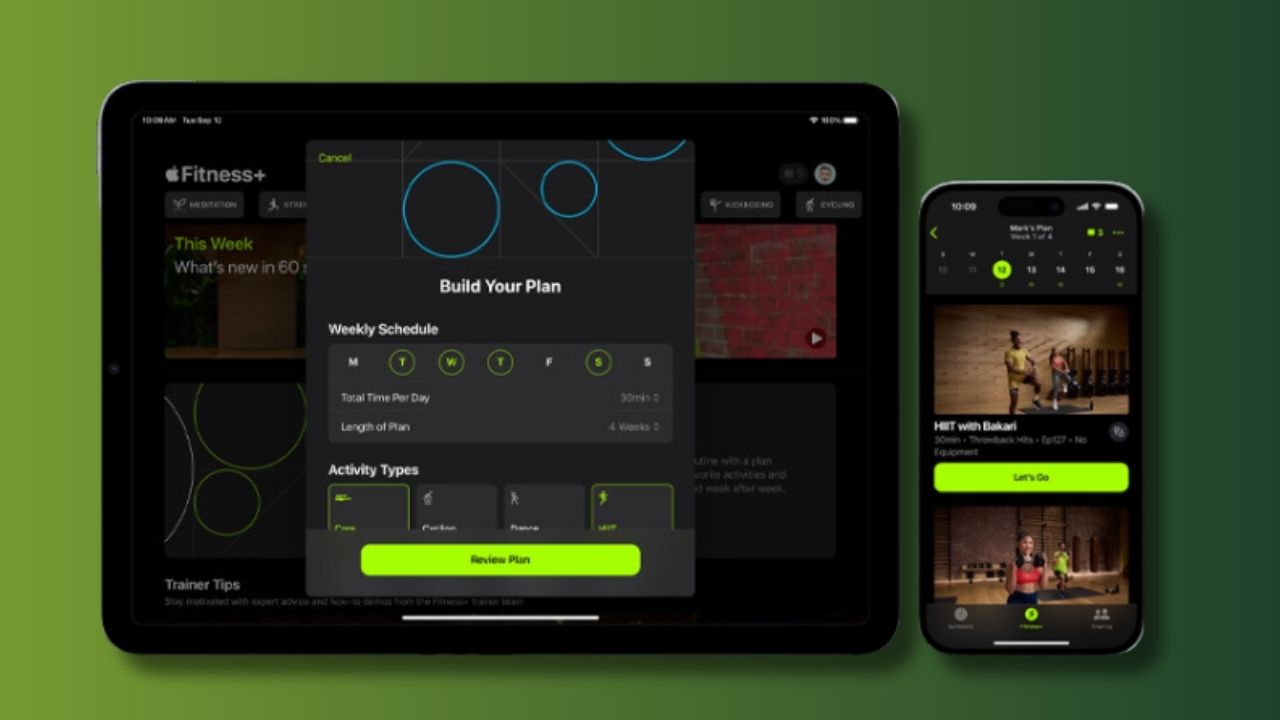

- Image Credit: Apple, Inc.
Since the pandemic hit, people have preferred at-home workouts over gym memberships. Naturally, more options for home workouts have become available. One of the most popular apps for a quality home workout is Apple Fitness +.
For only $10 a month, Apple Fitness + will help you achieve your goals. You can choose from various workouts, up to 45 minutes long, taught by actual fitness professionals. Your results are tracked in the app, making it easy to stay consistent and track your actions.
No more crowded gym floors or influencers hogging the equipment. Bring the gym to you or wherever you travel with the Apple Fitness + app.
15. Tonal Mirror


- Image Credit: Tonal.
It is hard to replicate lifting heavy weights and bars as you would in a gym, but the Tonal Mirror resistance technology is as close as you can get. The all-in-one workout machine comes with a wall-mounted screen that provides personalized coaching and fitness tracking. The equipment can hit all muscle groups and provide lifts like bench presses, squats, curls, and deadlifts.
The device can be a little pricey. At $3,000, it is a commitment, but over the course of a few years, the money saved on gym fees will pay for itself.