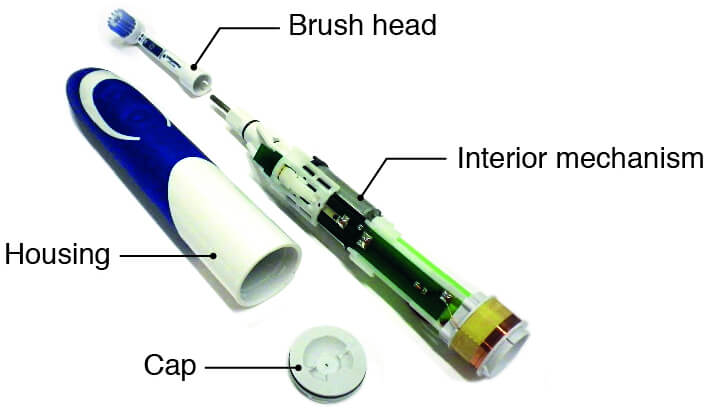
Nevadent NZAOT 600 A1 Premium Bluetooth elektromos fogkefe 8 pótfejjel, tokkal, időzítővel - eMAG.hu

Kedvezmény 4db/Csomag Csere Fogkefe Fej Szóbeli B Forgó Elektromos Fogkefe Puha Sörték Szóbeli Tisztítás Érdekel, Fogkefe Fej Szóbeli csere eredeti b > Áruház | Kereskedelmi-Eredeti.cam

Nevadent NZAOT 600 A1 Premium Bluetooth elektromos fogkefe 8 pótfejjel, tokkal, időzítővel - eMAG.hu

Vásárlás 4db elektromos fogkefe fej kefe fej csere orális higiénia b érzékeny ebs-17a a családi egészségügyi használata \ Outlet / www.clixxshoes.com

Vásárlás 4db elektromos fogkefe fej kefe fej csere orális higiénia b érzékeny ebs-17a a családi egészségügyi használata \ Outlet / www.clixxshoes.com

4DB Csere Elektromos Kefe Fej Oral-B A Gyerekek EB-10A Pro-Egészségügyi Szakaszában Gyermek Fogkefe Fejét AU Áruház Kategóriában. Elektromos Fogkefék And Csere Fej. Brandslux.news
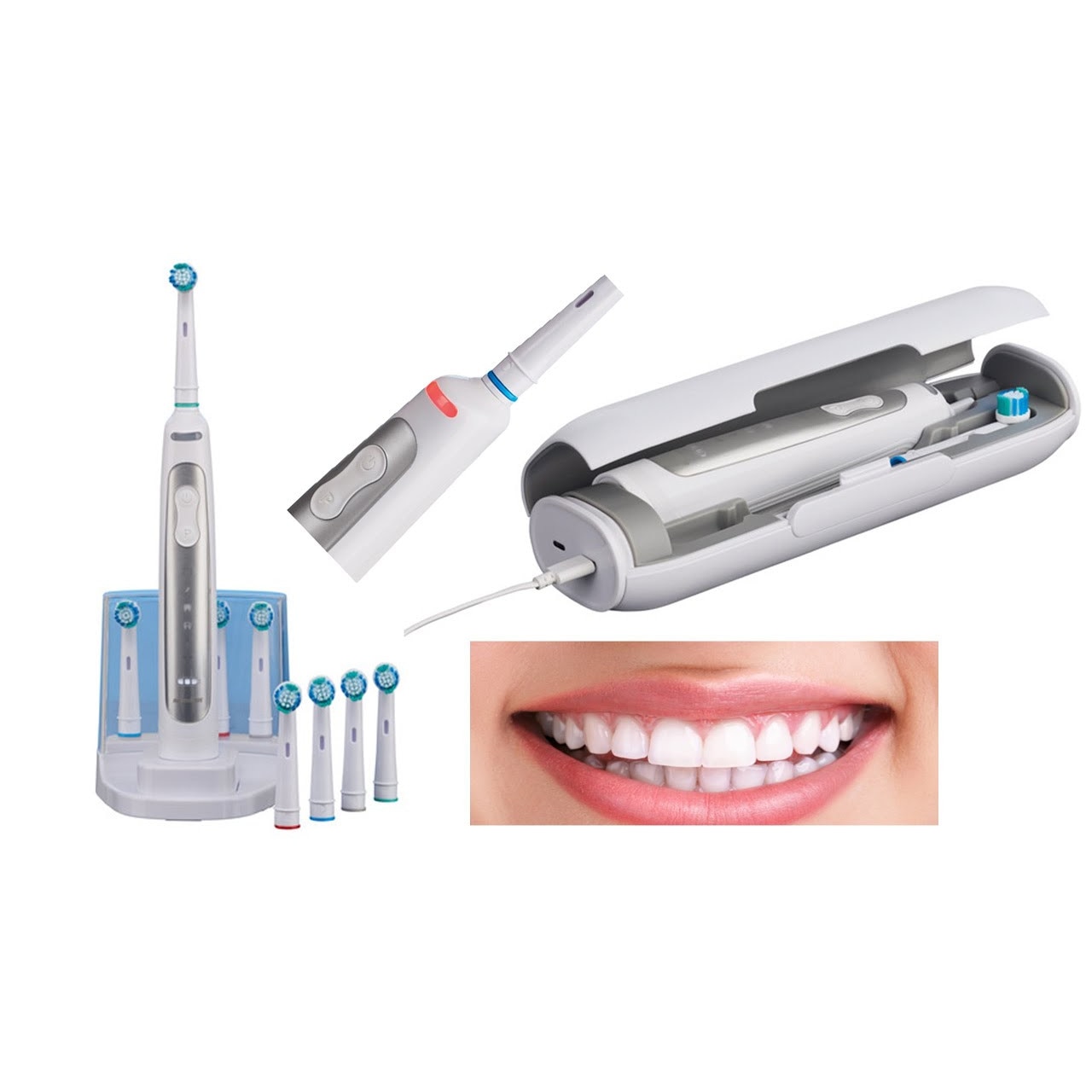
Nevadent NZAOT 600 A1 Premium Bluetooth elektromos fogkefe 8 pótfejjel, tokkal, időzítővel - eMAG.hu
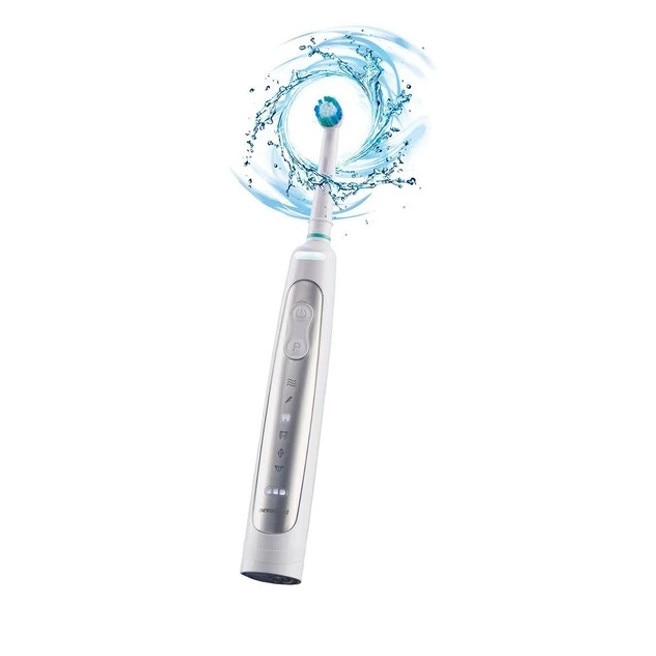
Nevadent NZAOT 600 A1 Premium Bluetooth elektromos fogkefe 8 pótfejjel, tokkal, időzítővel - eMAG.hu

Nevadent NZAOT 600 A1 Premium Bluetooth elektromos fogkefe 8 pótfejjel, tokkal, időzítővel - eMAG.hu

Vásárlás Arc Tisztító Kefe Fejét, 4db Csere Kefe Fej Oral-b Elektromos Fogkefe Illik Előre Power/pro-egészségügyi/diadal < outlet > www.vapordna.shop

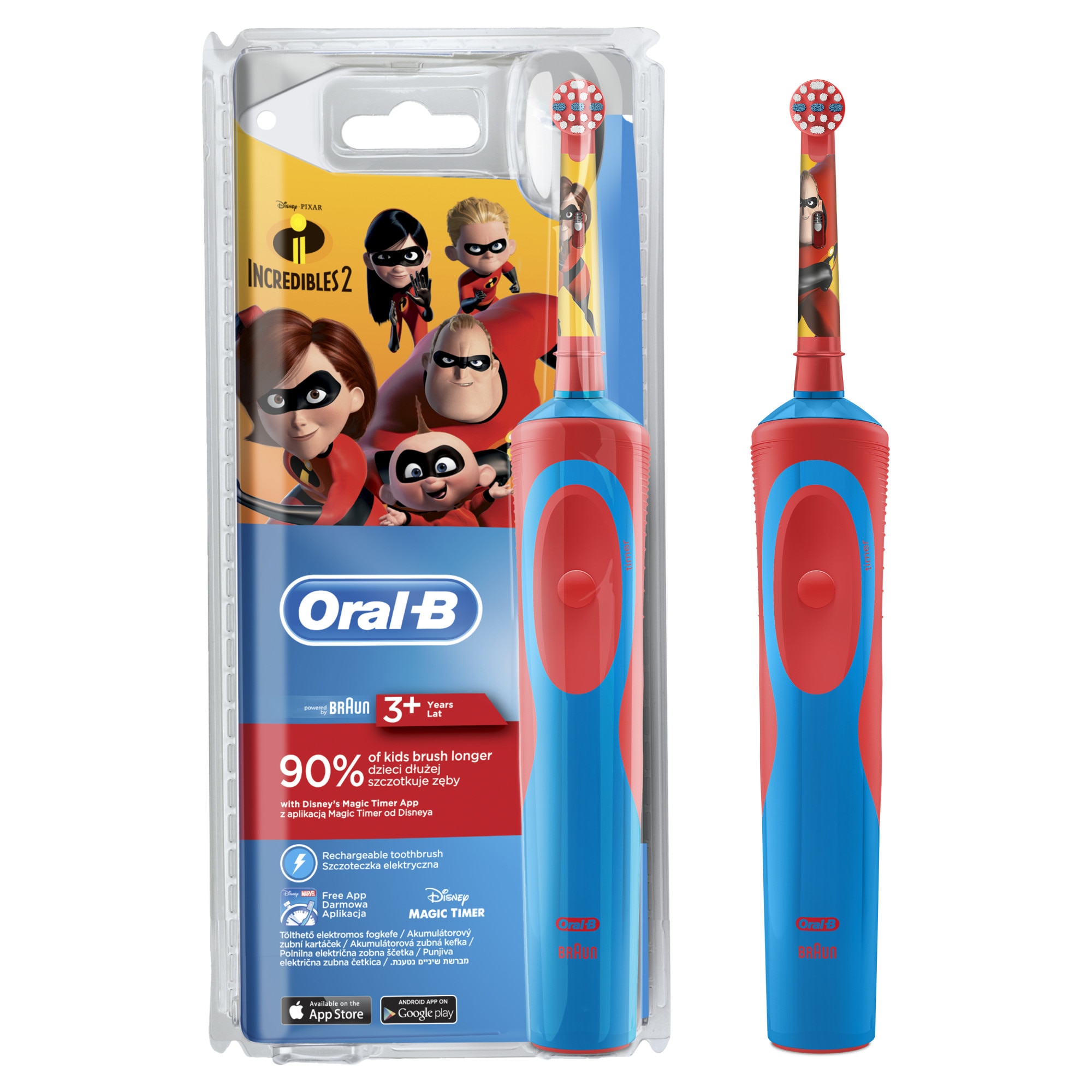
Oral b elektromos fogkefe kefe csere brushhead fúvóka + gyermekek csere fogkefe fej + védőborítás < A Legjobb > EgyetemesMarka.news
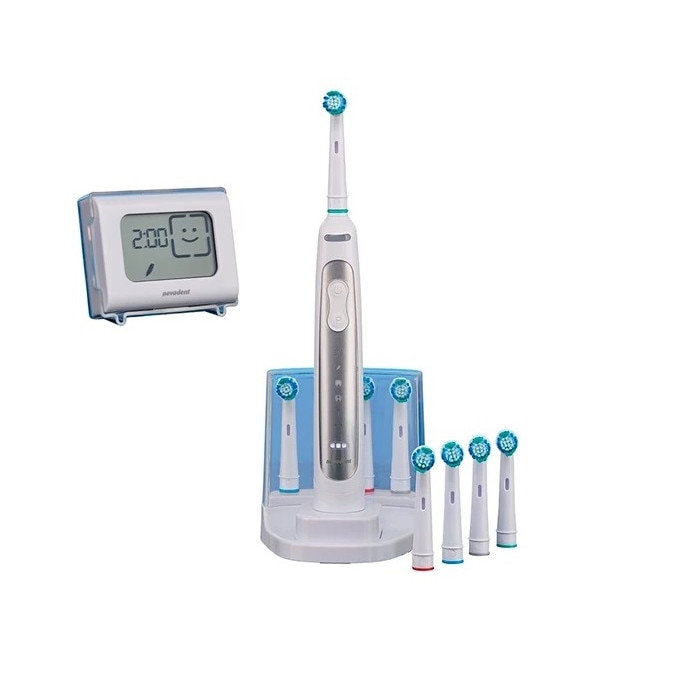
Nevadent NZAOT 600 A1 Premium Bluetooth elektromos fogkefe 8 pótfejjel, tokkal, időzítővel - eMAG.hu

Vásárlás 4db elektromos fogkefe fej kefe fej csere orális higiénia b érzékeny ebs-17a a családi egészségügyi használata \ Outlet / www.clixxshoes.com

Szónikus Elektromos Fogkefe Gyermekek Számára, A Gyerekek Ultrahangos Fogkefe Okos Szónikus Elektromos Fogkefe Hordozható Anti-slip Fogkefe > A legjobb - Alku-Konnyen.today