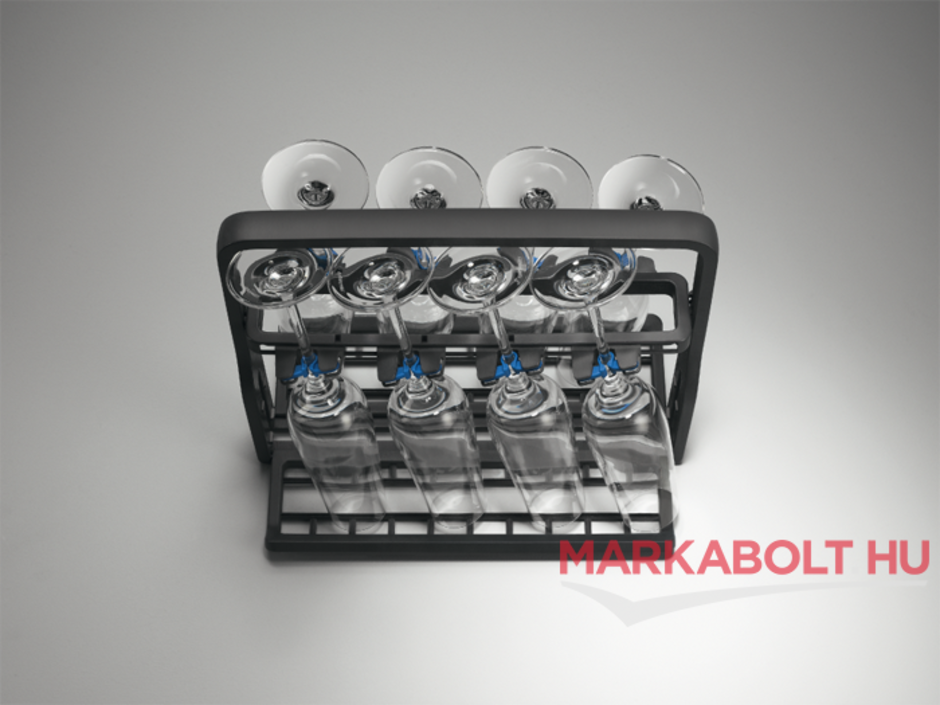
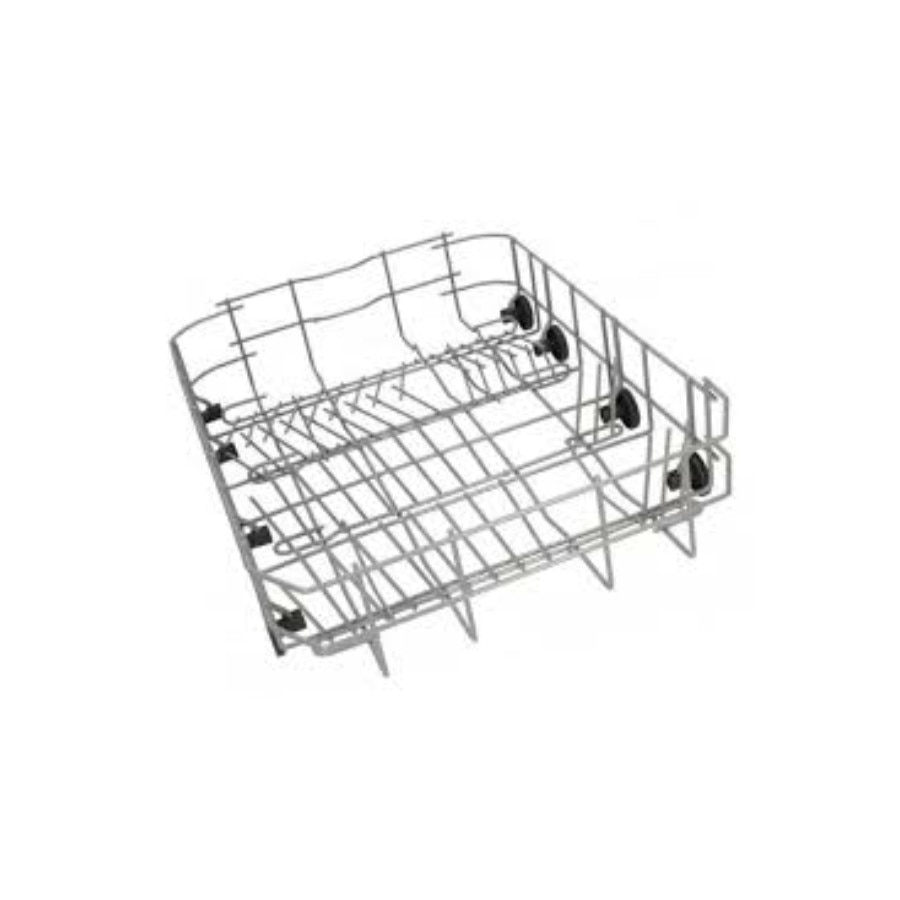
Electrolux pohártartó kosár :: Mosogatógép tartozékok :: AEG, Electrolux, Zanussi márkabolt webáruház

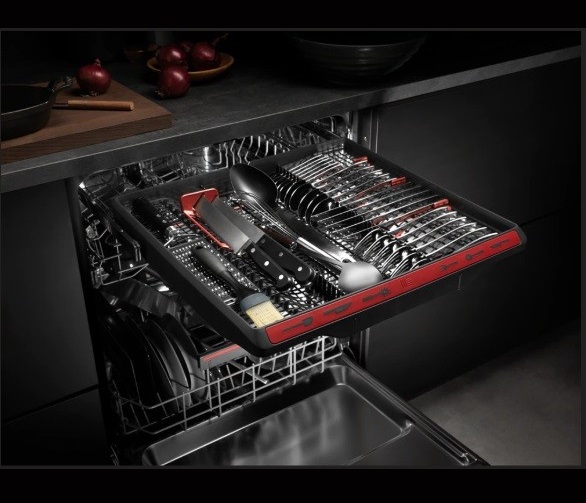
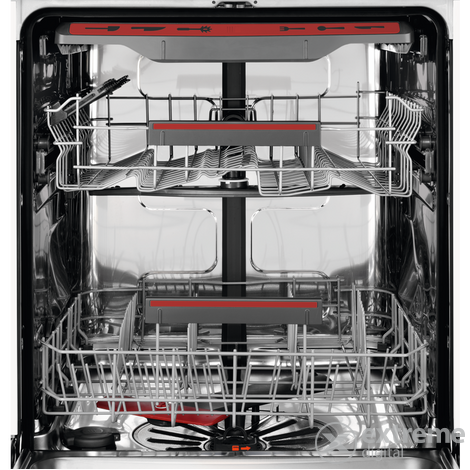
AEG FSB53927Z Beépíthető mosogatógép,14 teríték, QuickSelect kezelőpanel, MaxiFlex fiók, AirDry, 7 program

Electrolux pohártartó kosár :: Mosogatógép tartozékok :: AEG, Electrolux, Zanussi márkabolt webáruház
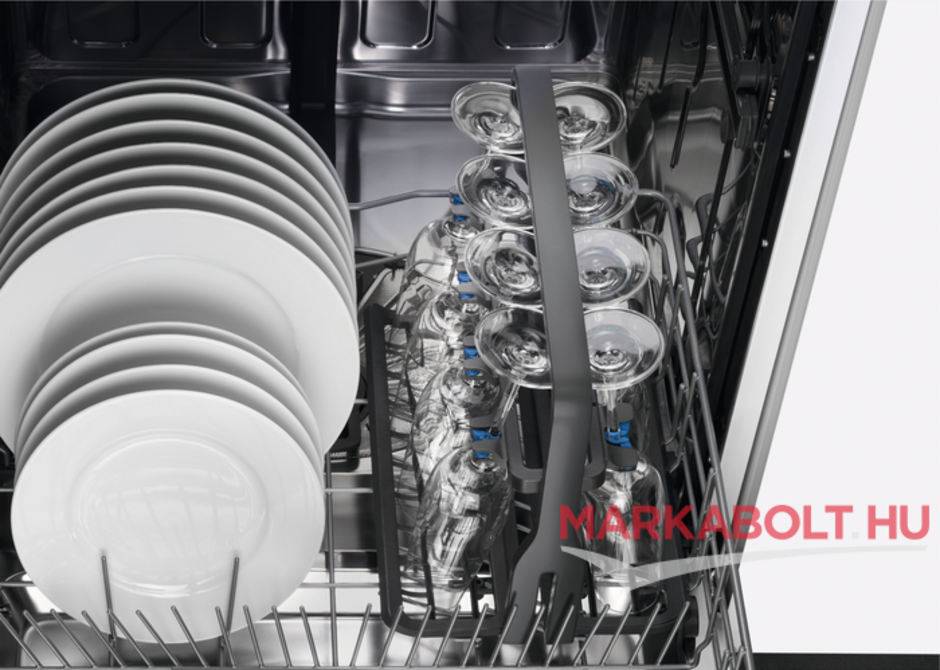

Electrolux pohártartó kosár :: Mosogatógép tartozékok :: AEG, Electrolux, Zanussi márkabolt webáruház
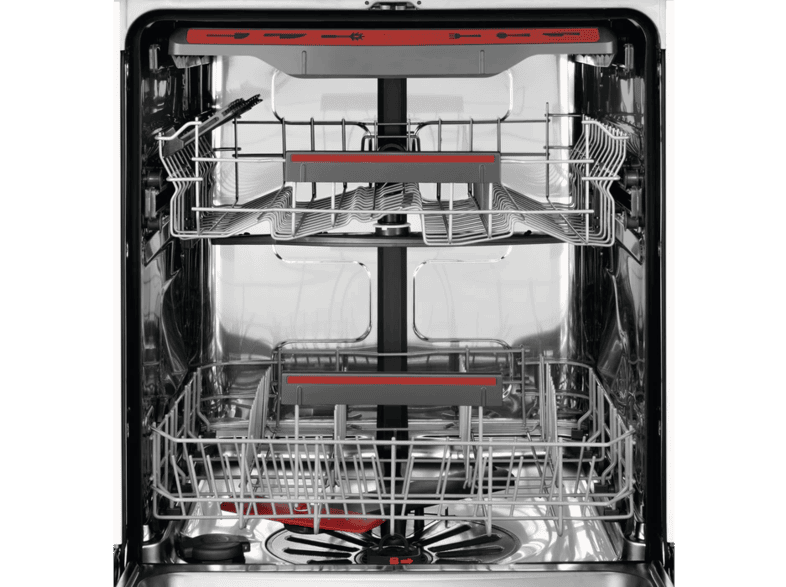

AEG FSB53927Z beépíthető mosogatógép, QuickSelect kezelőpanel, MaxiFlex fiók, 14 teríték, AirDry, 7 pr., 4 hőm. - MediaMarkt online vásárlás