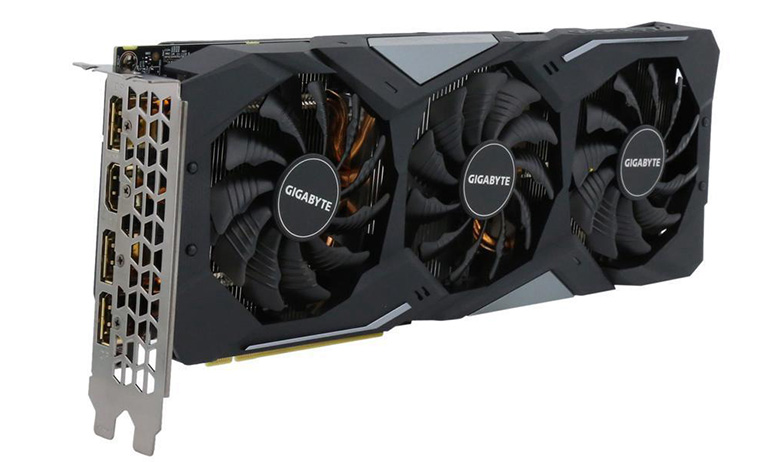
Gigabyte GTX 1660 SUPER GAMING OC review: Top 1660 SUPER capable of leapfrogging challenges | GearBest Blog
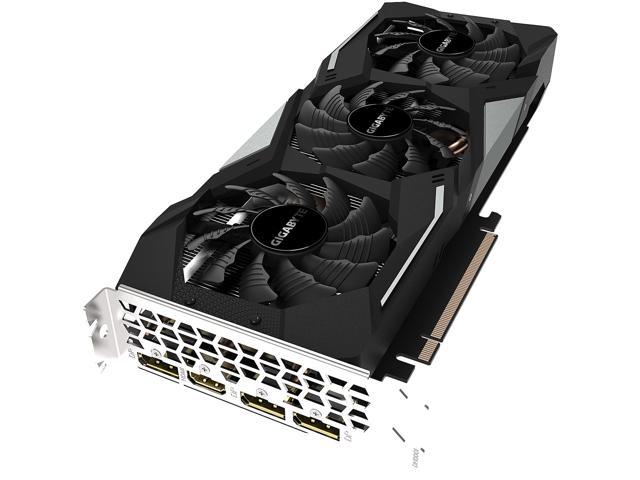

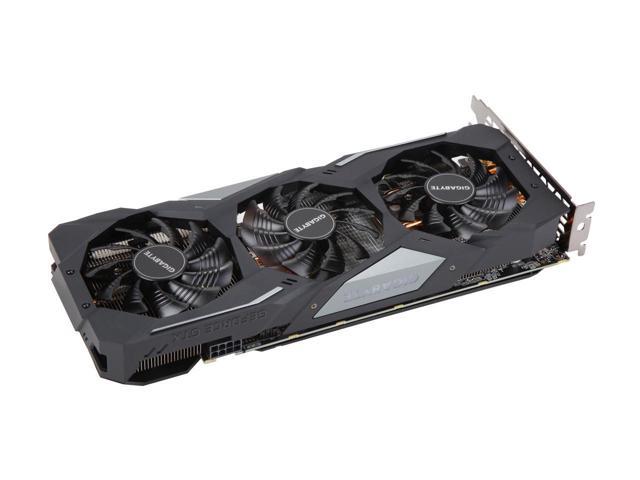
GIGABYTE GeForce GTX 1660 Ti GAMING OC 6G Graphics Card, 3 x WINDFORCE Fans, 6GB 192-Bit GDDR6, GV-N166TGAMING OC-6GD Video Card - Newegg.com

GIGABYTE GeForce GTX 1660 GAMING OC 6G Graphics Card, 3 x WINDFORCE Fans, 6GB 192-Bit GDDR5, GV-N1660GAMING OC-6GD Video Card - Newegg.com

Best Buy: GIGABYTE NVIDIA GeForce GTX 1660 OC Edition 6GB GDDR5 PCI Express 3.0 Graphics Card Black GV-N1660OC-6GD