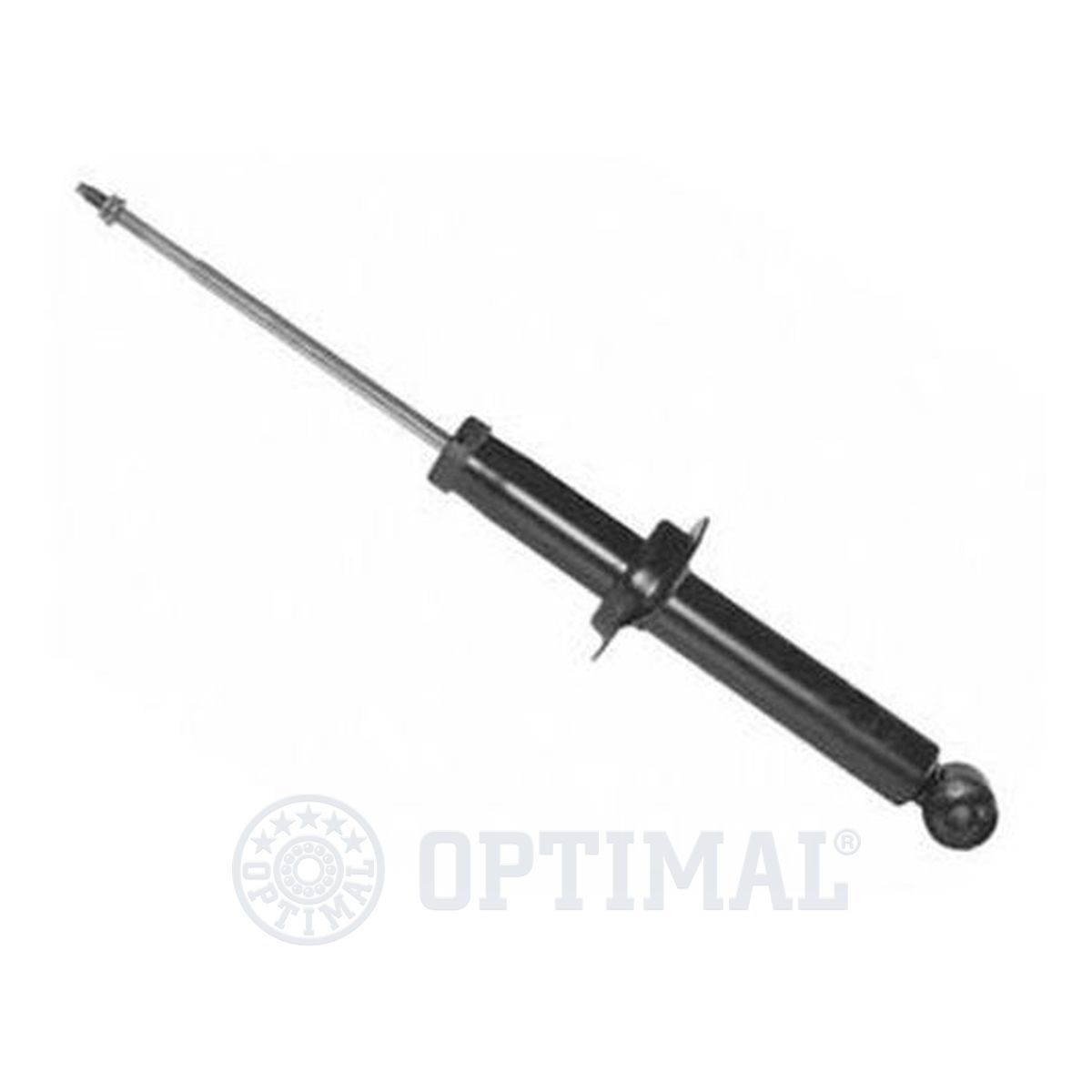
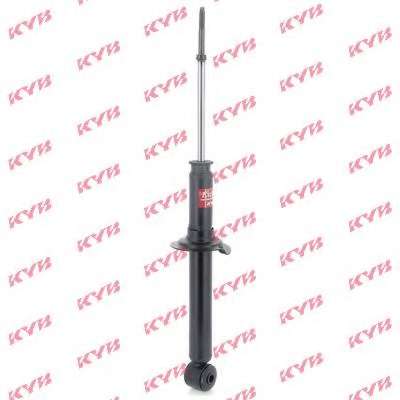
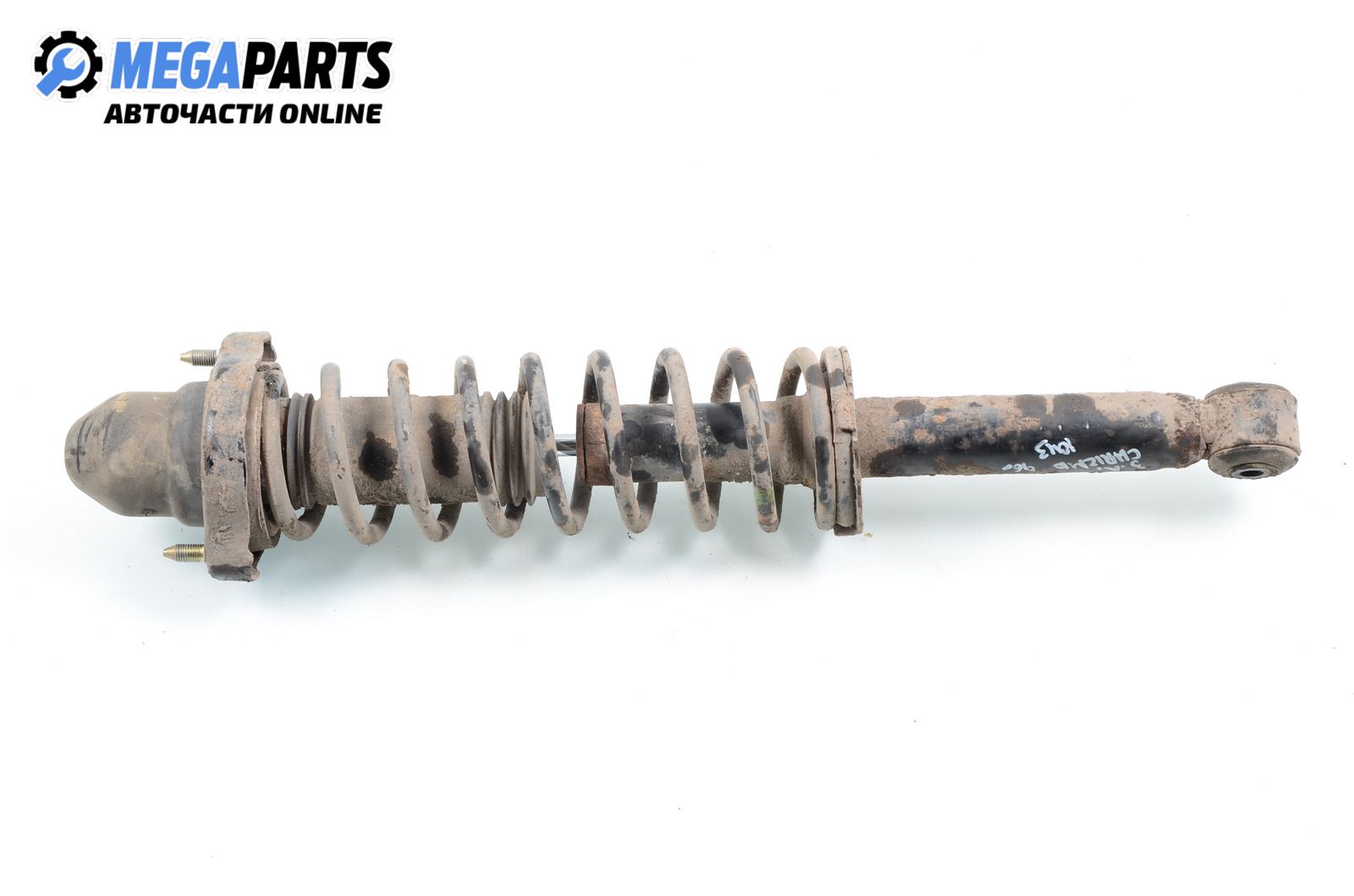
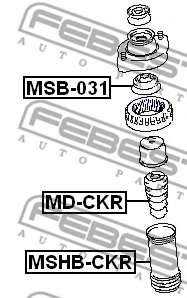
АМОРТИСЬОР за MITSUBISHI SPACE STAR » Амортисьор / спирална пружина / ресор Оригинални резервни части EU-АВТОЧАСТИ

Ударен тампон с прахоуловител на преден амортисьор за Mitsubishi Carisma Space Star . | Space stars, Pepper grinder, Stuffed peppers
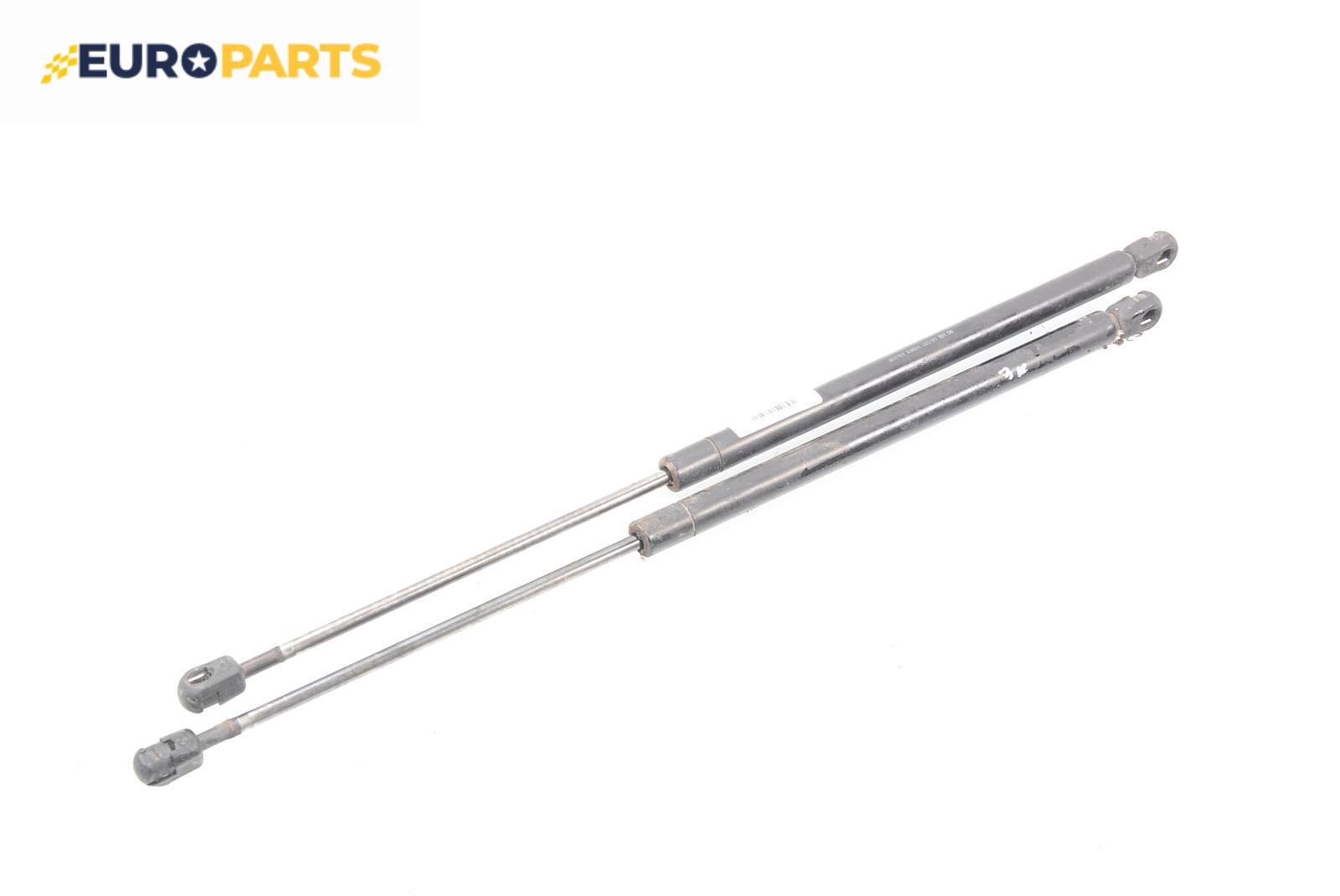

Амортисьор за багажник/товарно пространство за Mitsubishi Space Star Minivan (06.1998 - 12.2004) Цена: 10.70 лв. - EUROPARTS
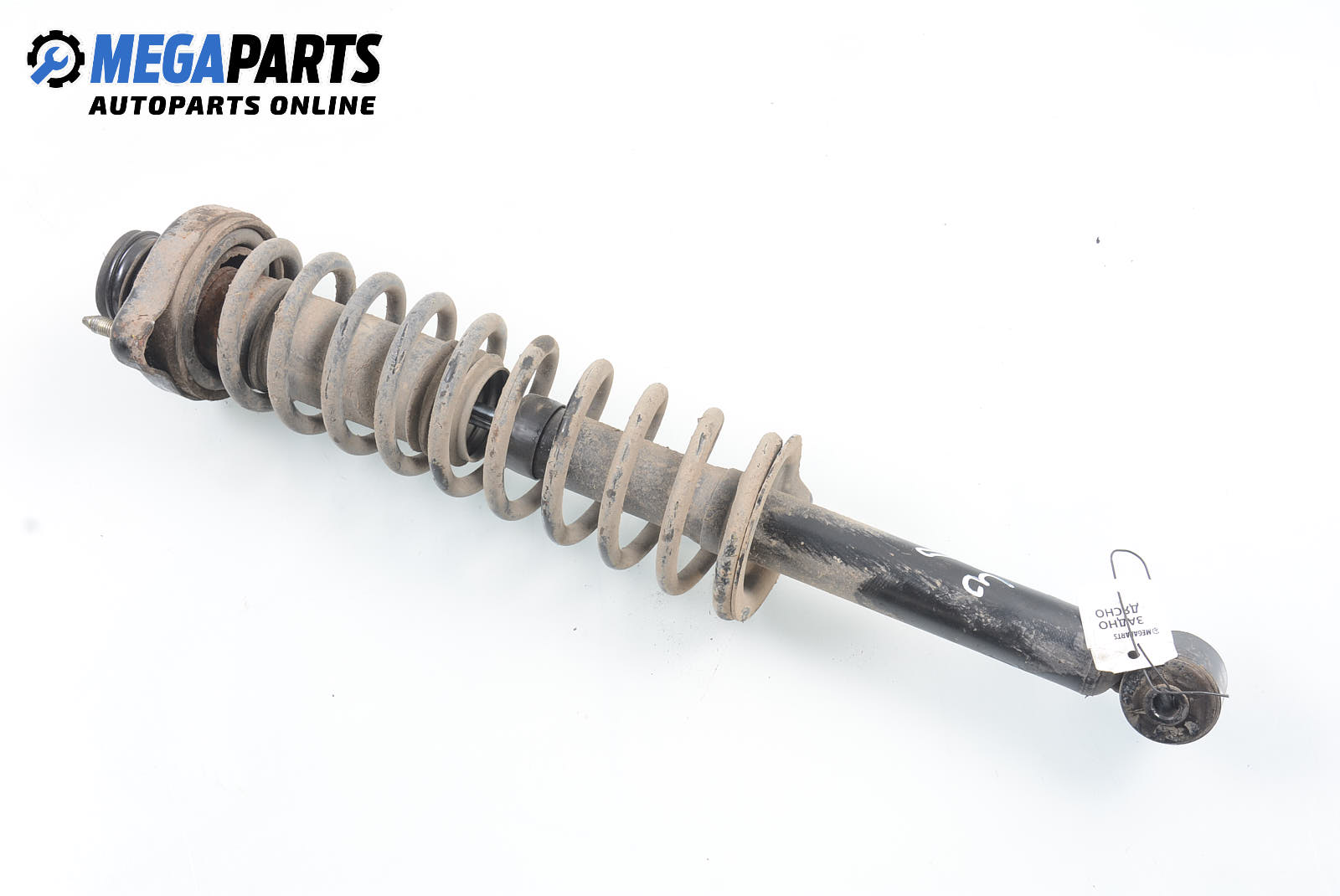

Макферсон за Mitsubishi Space Star Minivan (06.1998 - 12.2004), позиция: задна, дясна Цена: 38.70 лв. - MEGAPARTS
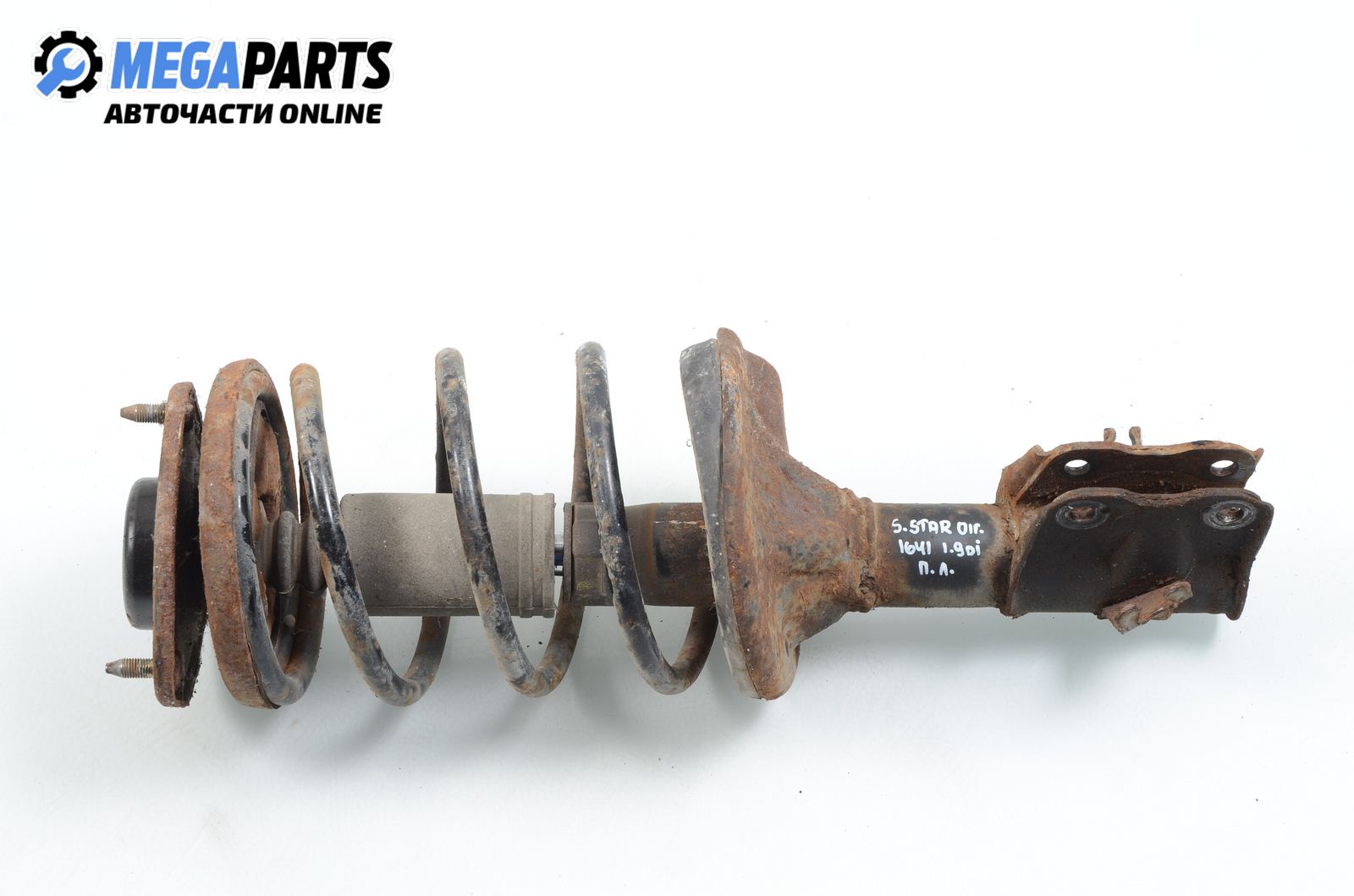

Макферсон за Mitsubishi Space Star Minivan (06.1998 - 12.2004), миниван, позиция: предна, лява Цена: 65.00 лв. - MEGAPARTS