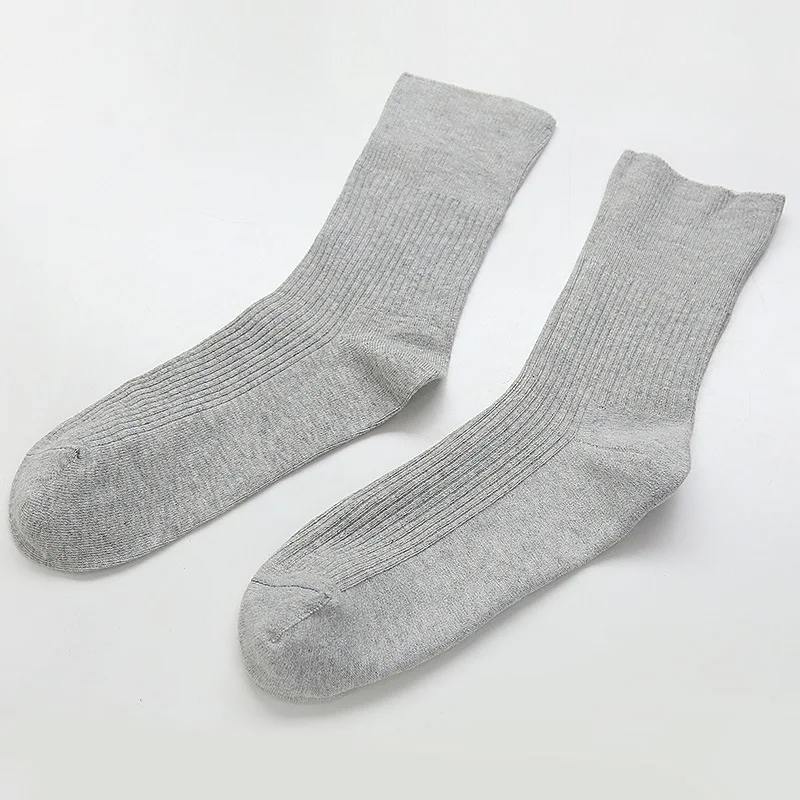
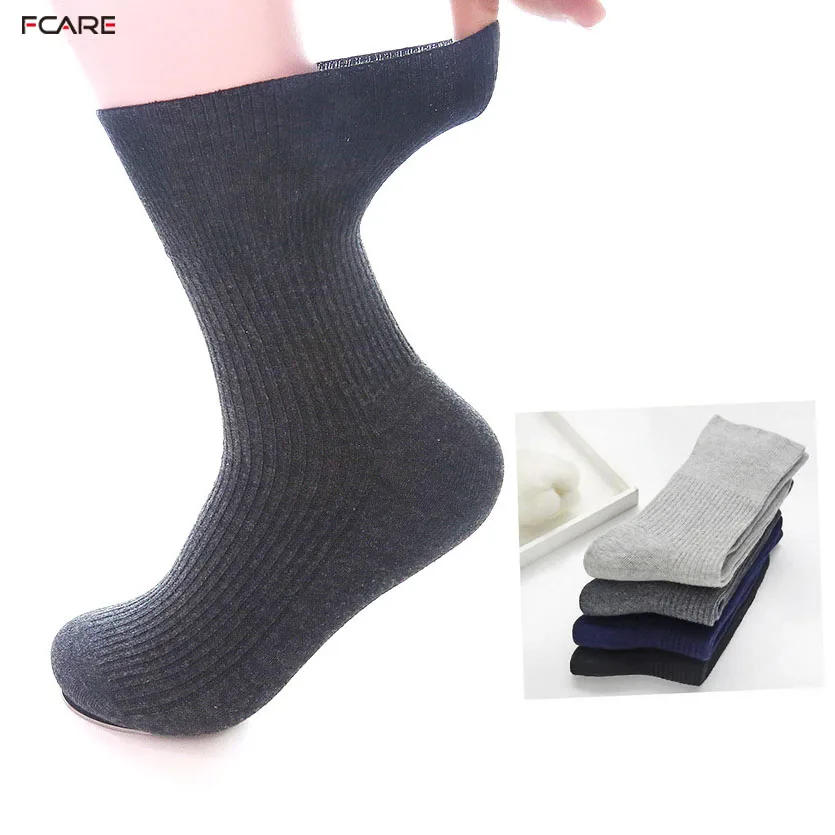
Fcare 10шт=5 двойки плюс размера на хипертония чорапи предотвратяване на разширени вени чорапи 44-48eu диабет есен-зима неизмита памучни чорапи купи онлайн > нов \ Sales-Original.cam
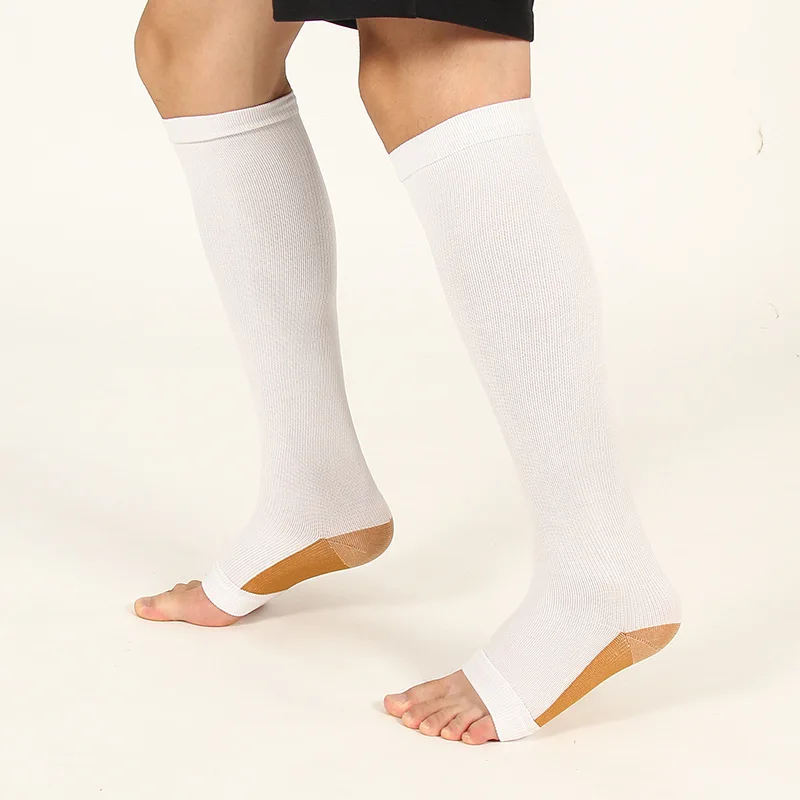

Купи онлайн Мъжете Компресия Чорапи Чорапи Пакет Унисекс Спортни Чорапи Много Предотвратяване На Разширени Вени Медицинска Сестра Чорапи Футбол Бягане ~ Бельо и Пижами - Energybg.eu

сантиметър практикуващ лекар радикален стягащи ластични чорапи за разширени вени - semeandoeducacao.org

Отстъпка Чорапи унисекс компресия чорапи за колоездене чорапи, подходящи за медицински сестри,лекари, учители, отоци, диабет, разширени вени чорапи \ Бельо и пижами > News365.news

Компресионни чорапи против разширени вени купи в интернет - магазин DREBISIMO: цени, снимки, характеристики
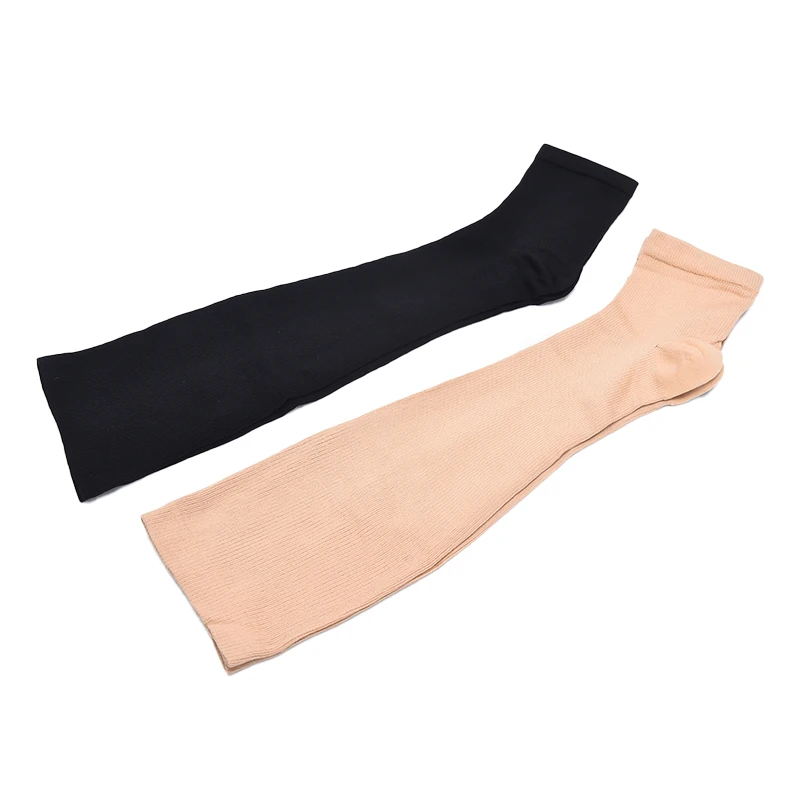

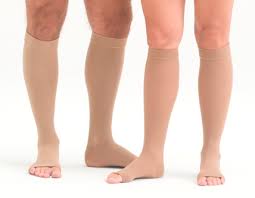
Еластичен Открит Чорап Коляното Чорапи Телешки Компресия Чорапи Разширени Вени, Лечение на Формирането на Степен Чорапи Налягане S-XL купи онлайн | Бельо и Пижами \ www.different-stylebg.com

Размер S-XXL Компресия Чорапи Медицинска Сестра Разширени Вени, Подуване на Диабет Кръвообращението Грижа Против Умора Компресия чорапи купи \ Бельо и пижами / Lopters.today

Купи Adult Keep Going Short Crew Street Fashion Памучни Чорапи Youth Daily Teen Human Streetwear Snail Logo Агитация Ободряющее Дума > Бельо и пижами | www.inkling.news

10 бр. дамски чорапи памучни сладки мультяшные жените чорапи бродирани забавни къси летни чорапи удобни ежедневни горещи носители Разпродажба! - Дамски чорапи и трикотаж-носочные на продукта < Bbangla.news

Поръчка 2 чифта/комплект дамски памучни чорапи Harajuku Casual Warm Long-sock Girls Multicolor Socken мода есен-зима сокс Streetwear #f / изход - Revelnail.shop
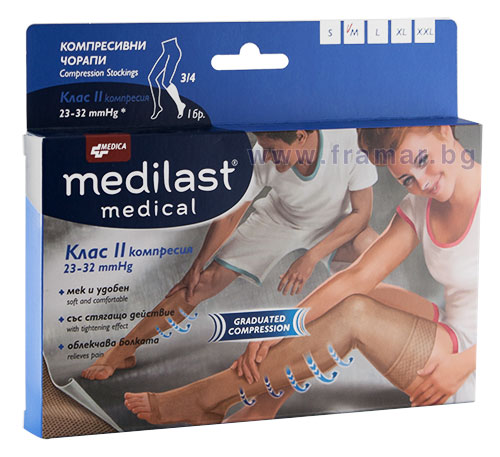
сантиметър практикуващ лекар радикален стягащи ластични чорапи за разширени вени - semeandoeducacao.org
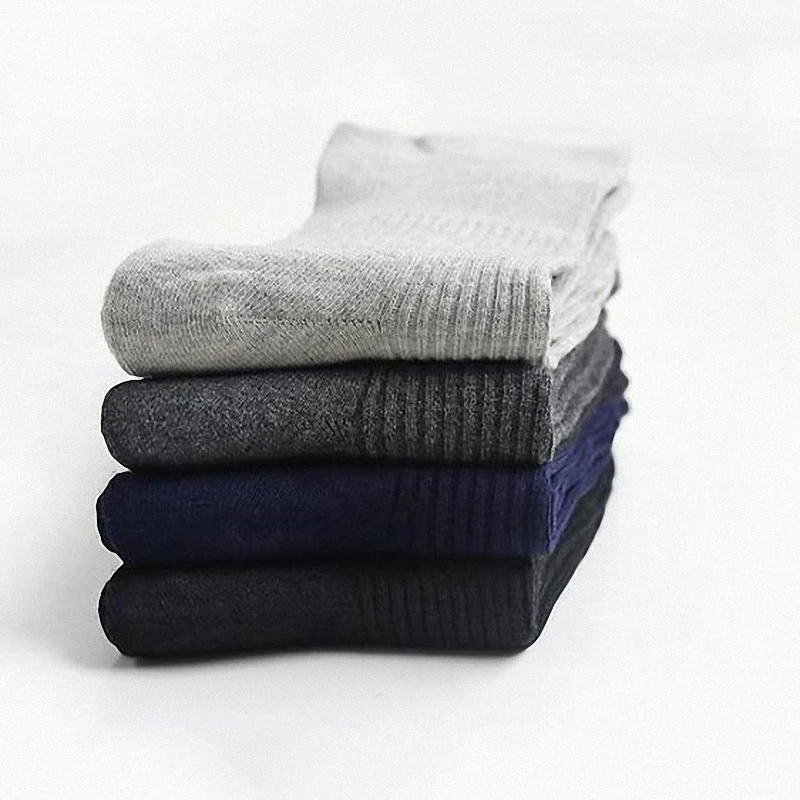

Fcare 10шт=5 двойки плюс размера на хипертония чорапи предотвратяване на разширени вени чорапи 44-48eu диабет есен-зима неизмита памучни чорапи купи онлайн > нов \ Sales-Original.cam

Купи Разширени Вени, Болки В Краката Чорапогащи Налягане На Компресия Чорапи Унисекс Цвят На Бедрата Високи Чорапи, Найлонови Чорапи, Дълги Чорапи Нови < Дамски чорапи и трикотаж-носочные на продукта - Geekculture.news

Еластичен Открит Чорап Коляното Чорапи Телешки Компресия Чорапи Разширени Вени, Лечение на Формирането на Степен Чорапи Налягане S-XL купи онлайн | Бельо и Пижами \ www.different-stylebg.com
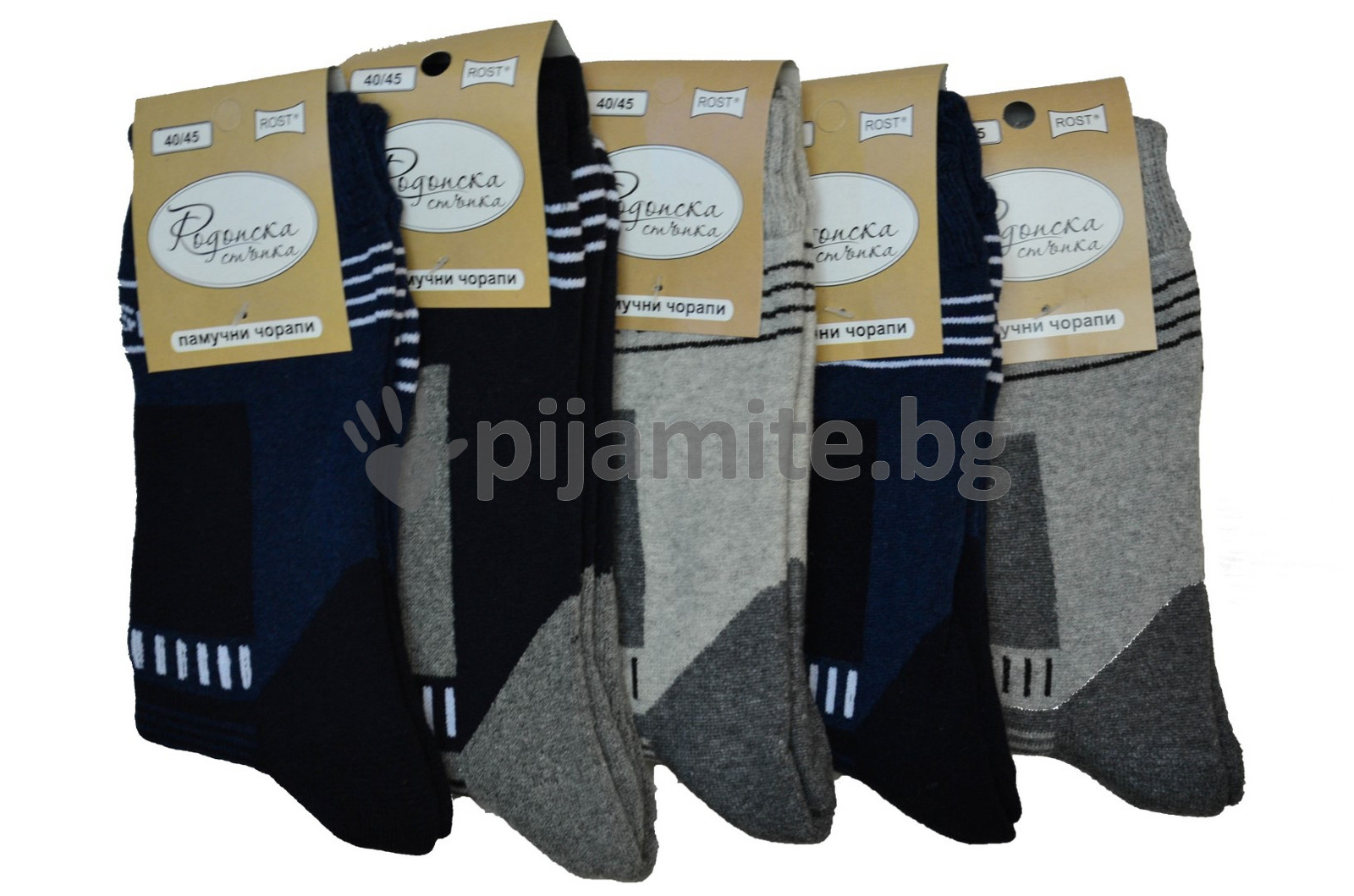

Мъжки Термо чорапи свободен ластик, разширени вени 40/45 - 5 бр./пакет, , | Български пижами и бельо

Купи Разширени Вени, Болки В Краката Чорапогащи Налягане На Компресия Чорапи Унисекс Цвят На Бедрата Високи Чорапи, Найлонови Чорапи, Дълги Чорапи Нови < Дамски чорапи и трикотаж-носочные на продукта - Geekculture.news

Купи Разширени Вени, Болки В Краката Чорапогащи Налягане На Компресия Чорапи Унисекс Цвят На Бедрата Високи Чорапи, Найлонови Чорапи, Дълги Чорапи Нови < Дамски чорапи и трикотаж-носочные на продукта - Geekculture.news