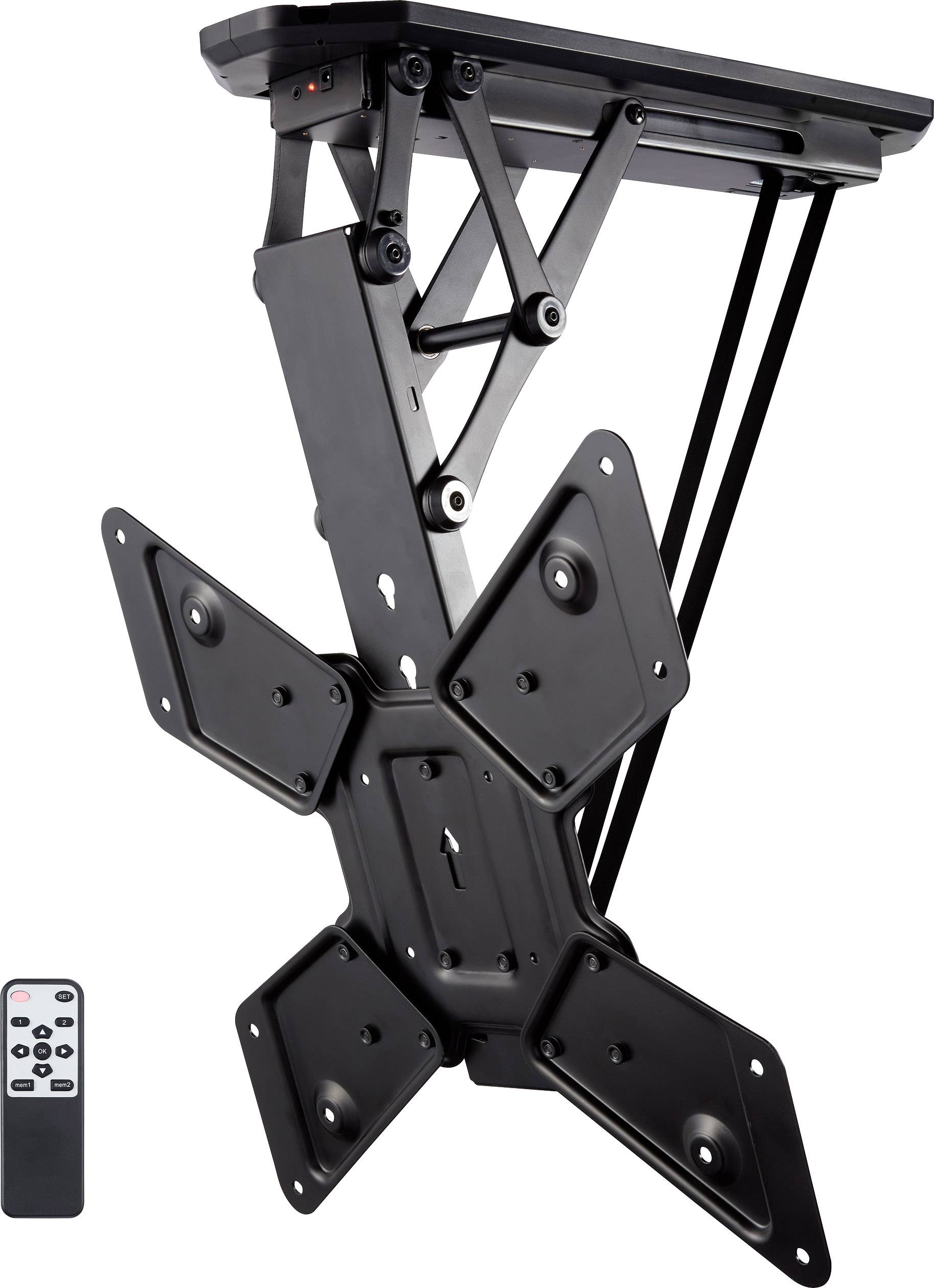
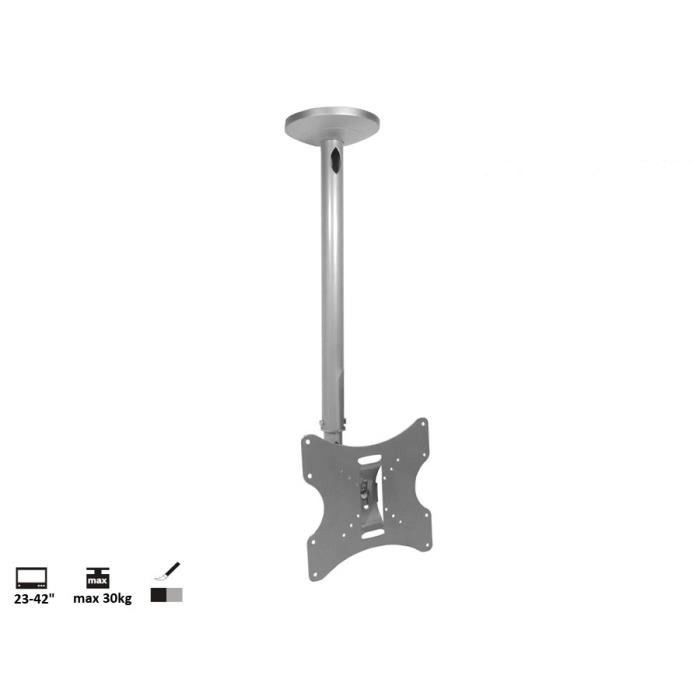
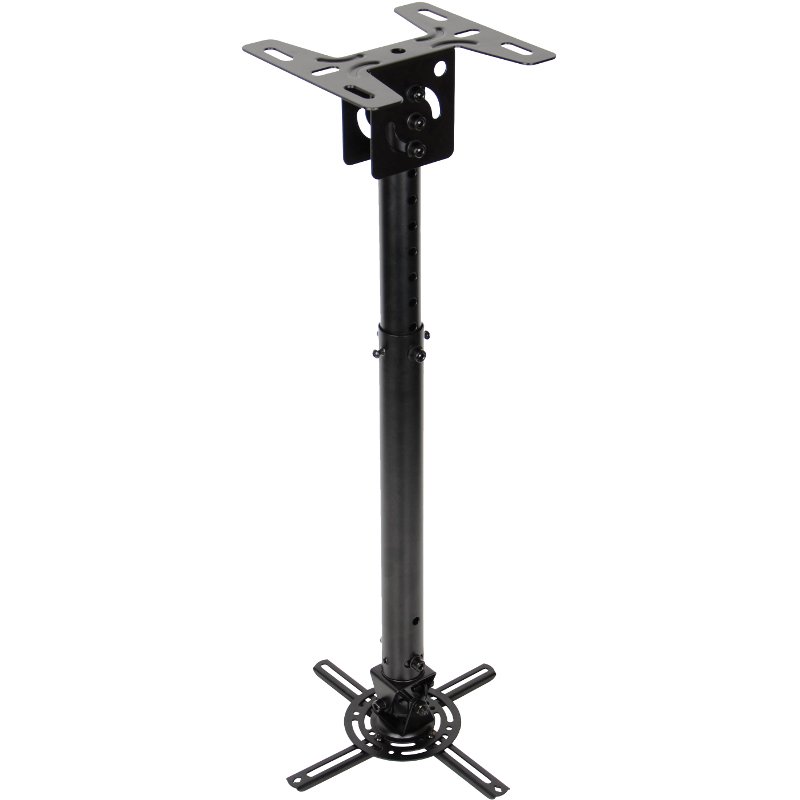
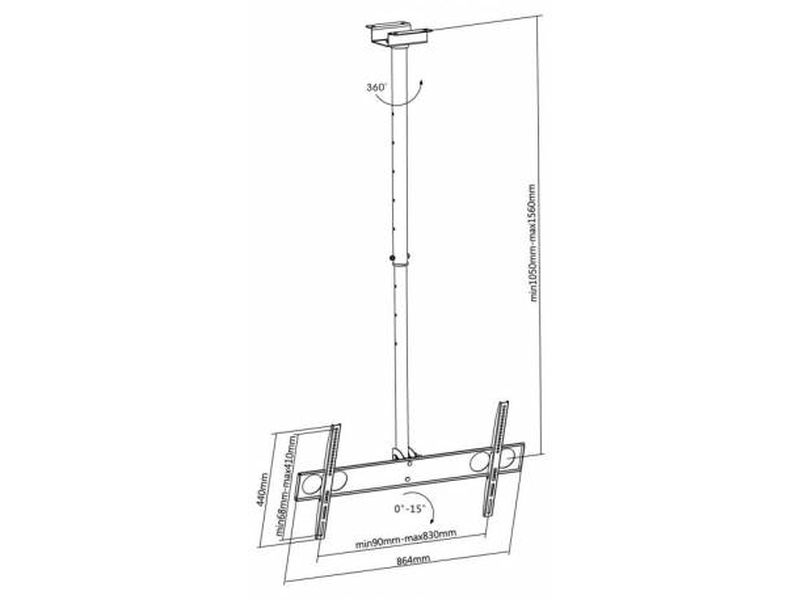
Motoros mennyezeti TV tartó konzol távirányítóval 58,4 - 139,7 cm 23 - 55" dönthető, SpeaKa Professional | Conrad

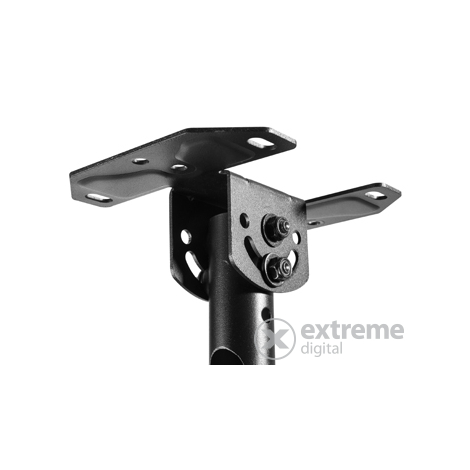
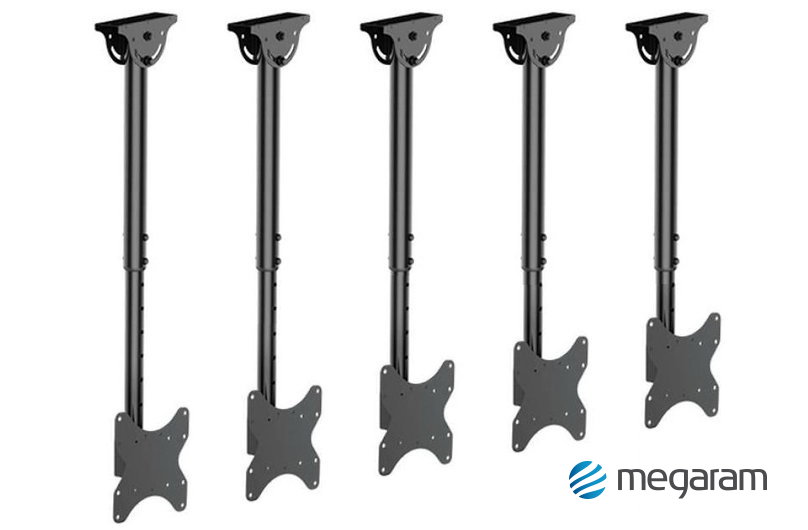
Rendelés 5db Univerzális TV Fali Konzol 360Degree Forgó Hangszóró Konzol Mennyezeti Fali Tartók Beltéri Kültéri Kamera Monitor Jogosultja / Otthoni elektronikai tartozékok > PiacterErtek.today
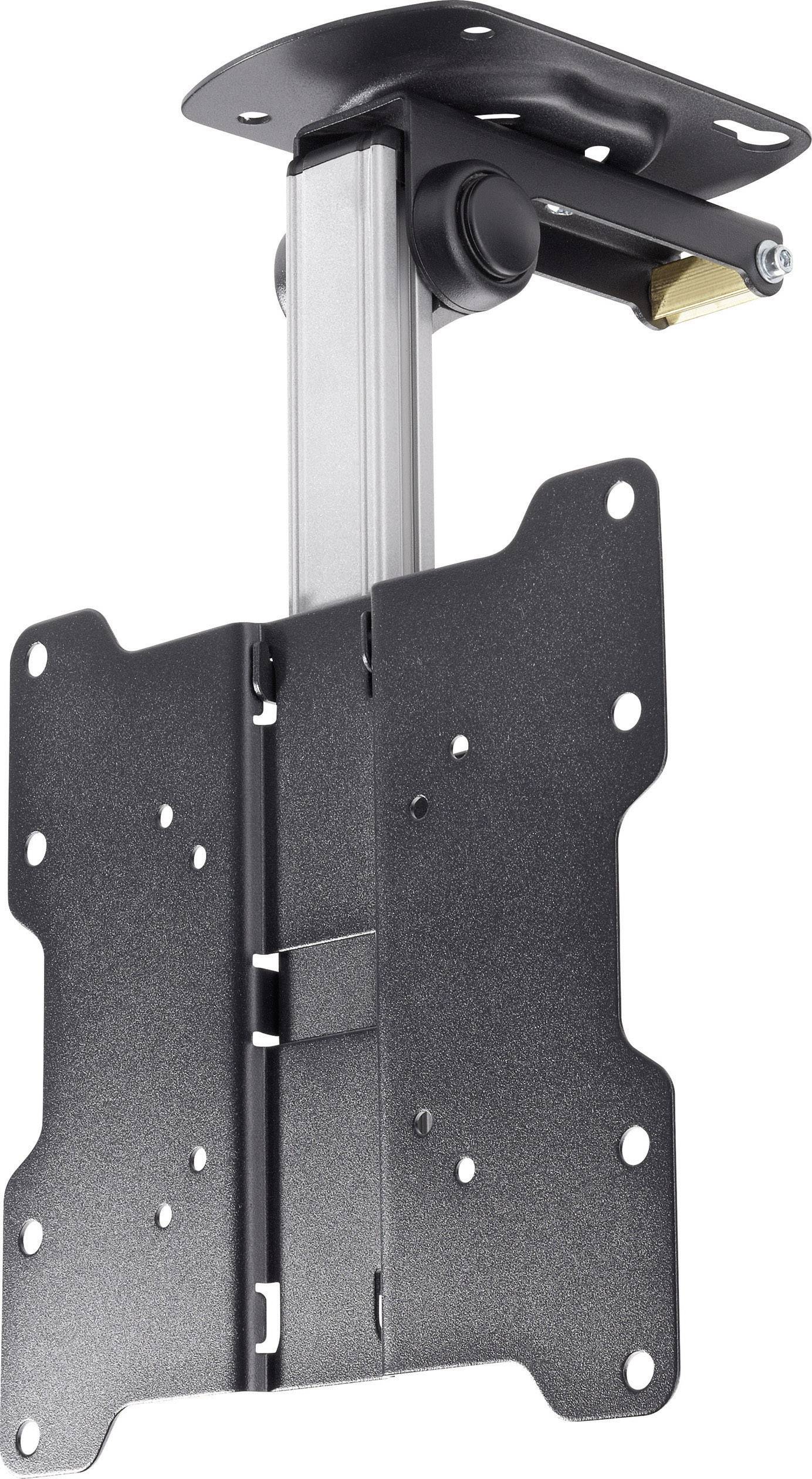

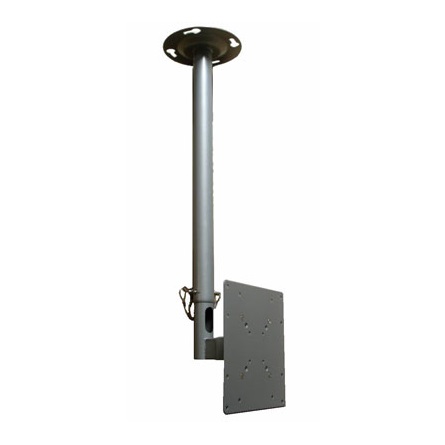
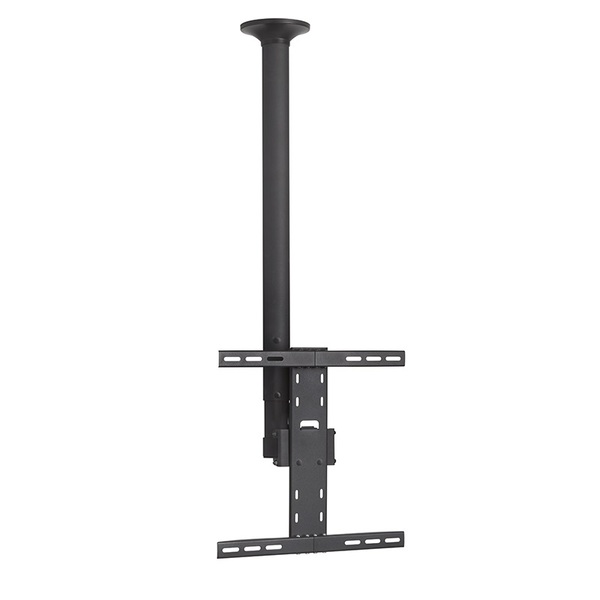
Mennyezeti TV tartó konzol 43 - 94 cm 17 - 37", dönthető, forgatható, SpeaKa Professional 629563 | Conrad