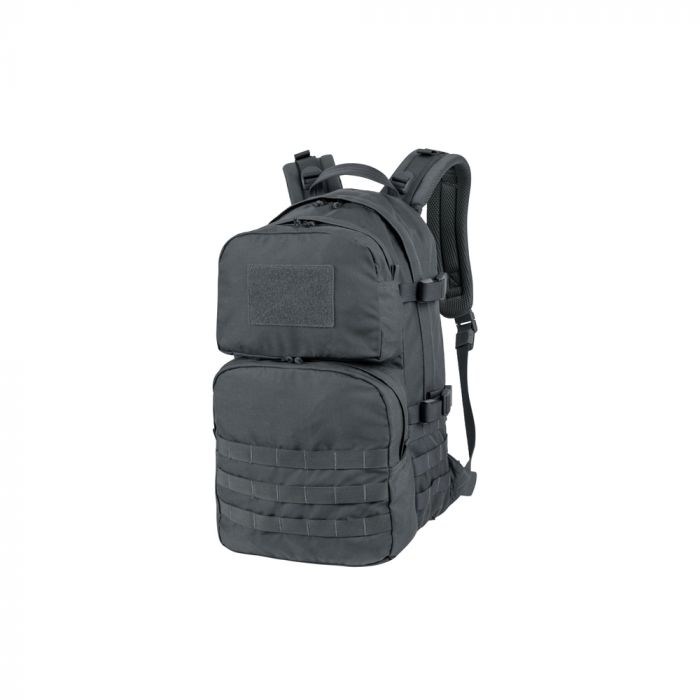
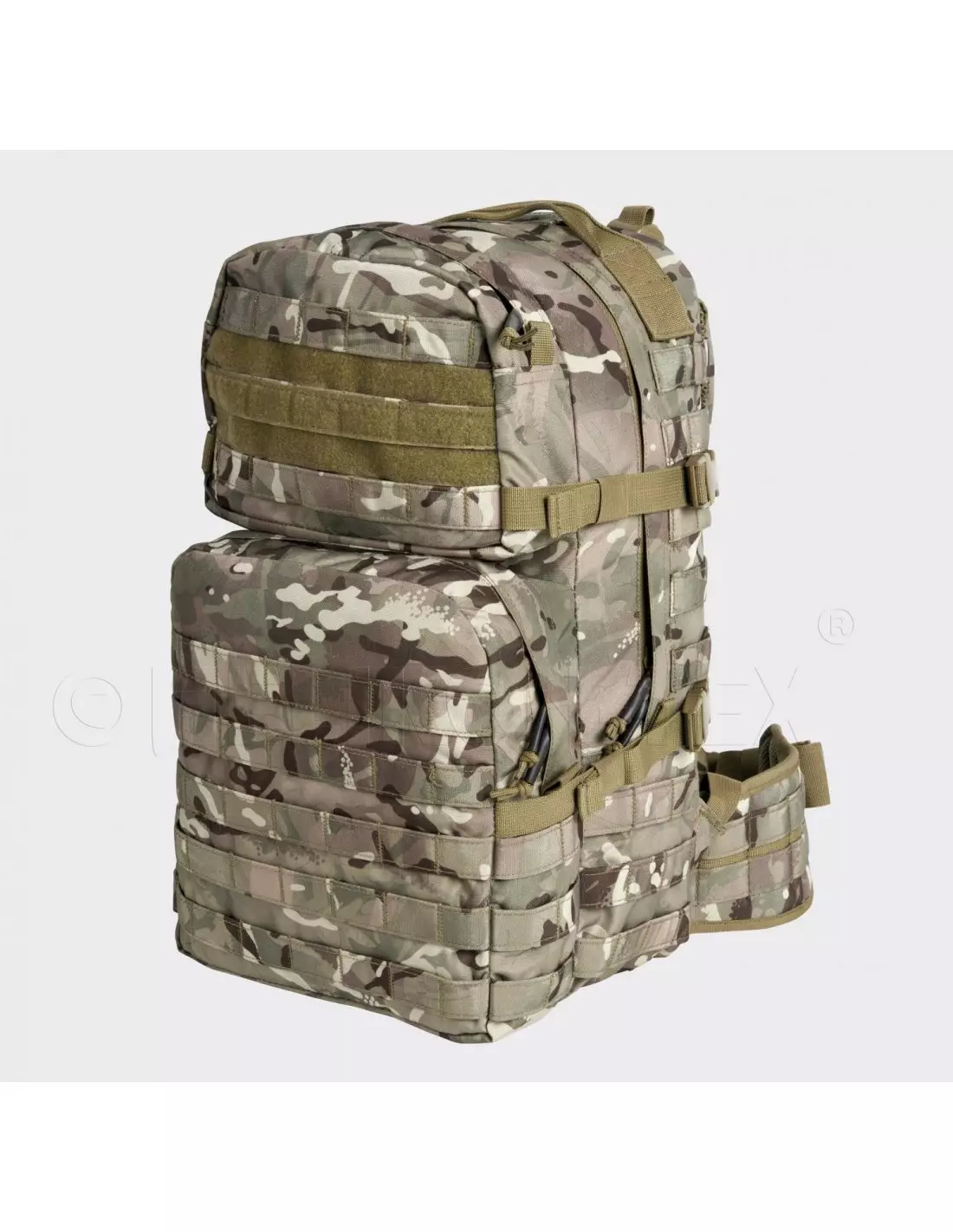
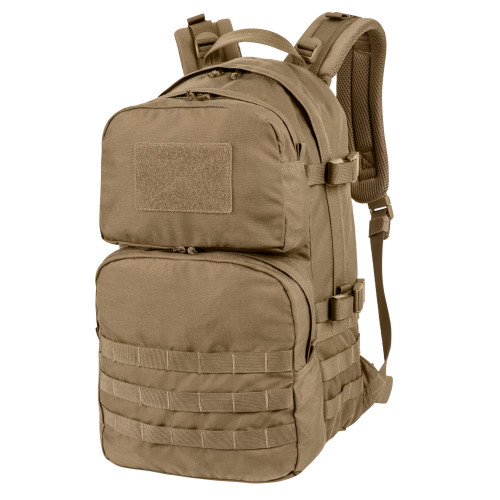
Helikon - Plecak Ratel Mk2 - 25 L - Pantera Leśna - PL-RT2-CD-04 - Plecaki - Plecaki i Torby - Wyposażenie taktyczne - airsoft, militaria, ASG - redberet.pl

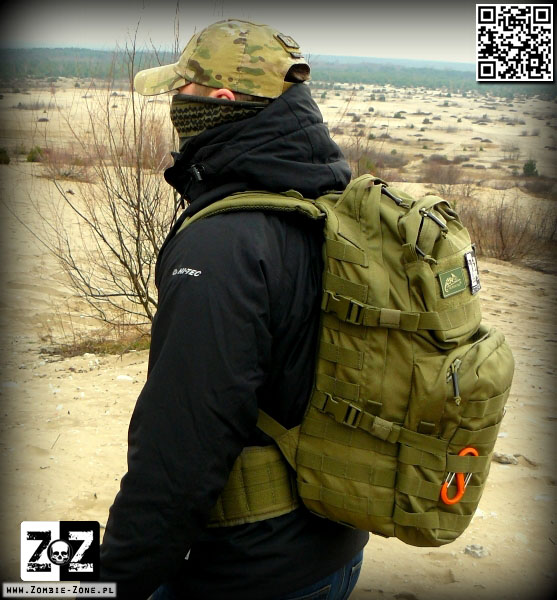
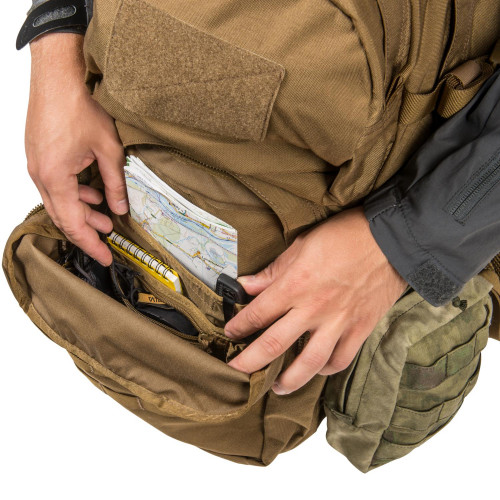
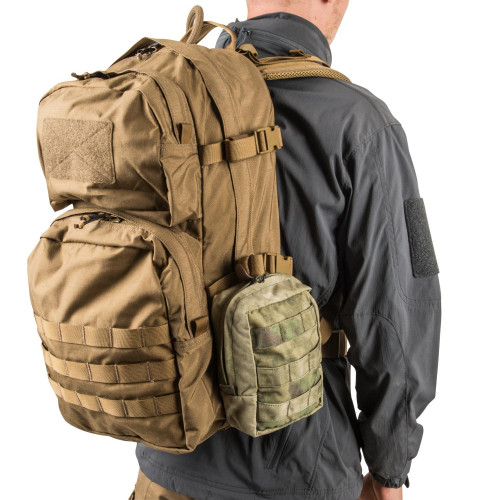
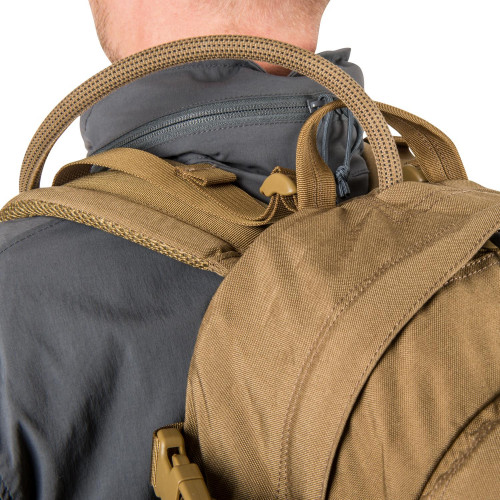
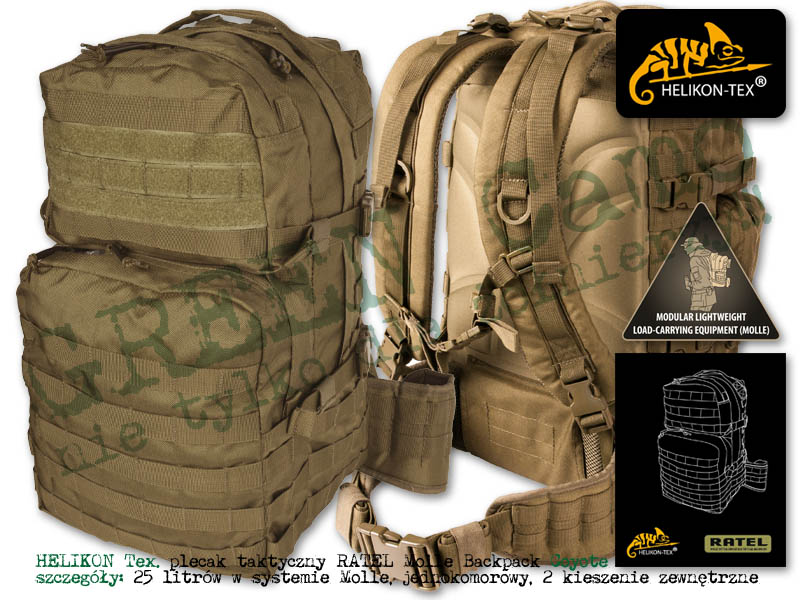
Plecak Helikon-Tex RATELSurvival, Zombie, Tactical, Outdoor – Zombie Zone | Survival, Zombie, Tactical, Outdoor - Zombie Zone
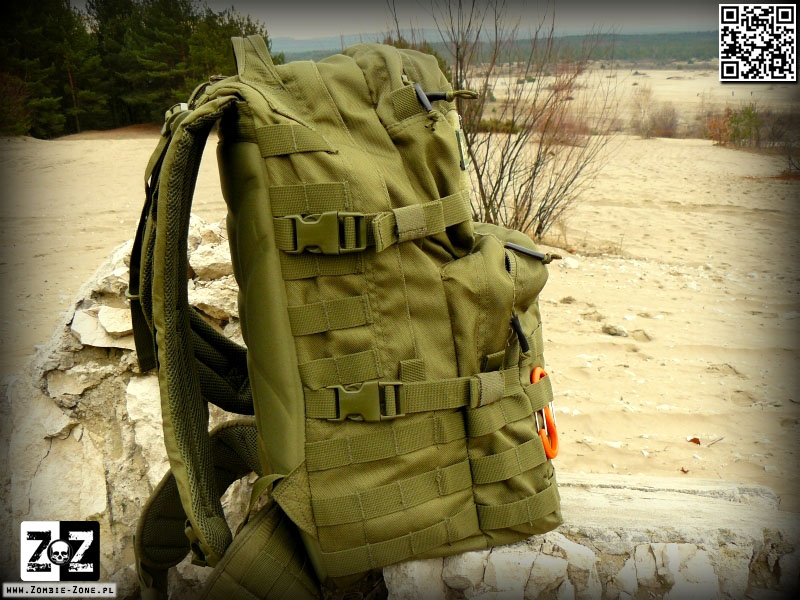

Plecak Helikon-Tex RATELSurvival, Zombie, Tactical, Outdoor – Zombie Zone | Survival, Zombie, Tactical, Outdoor - Zombie Zone