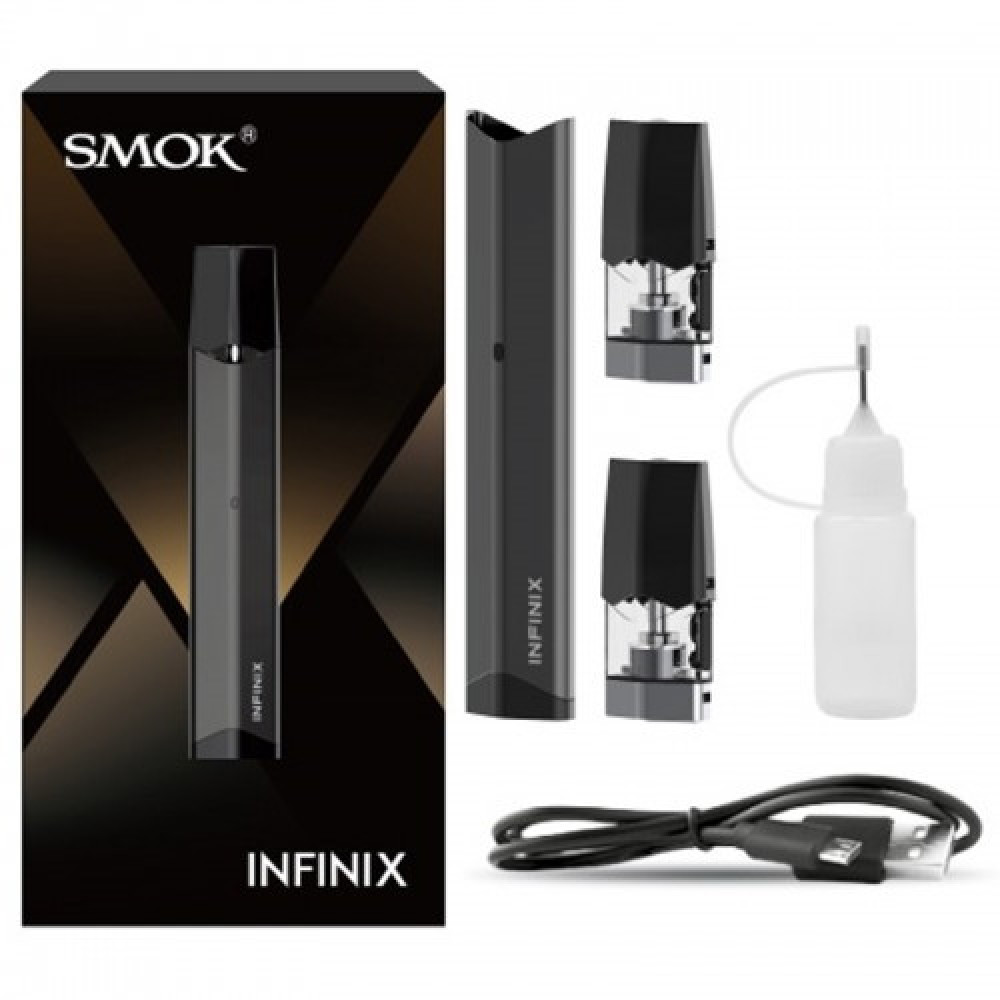
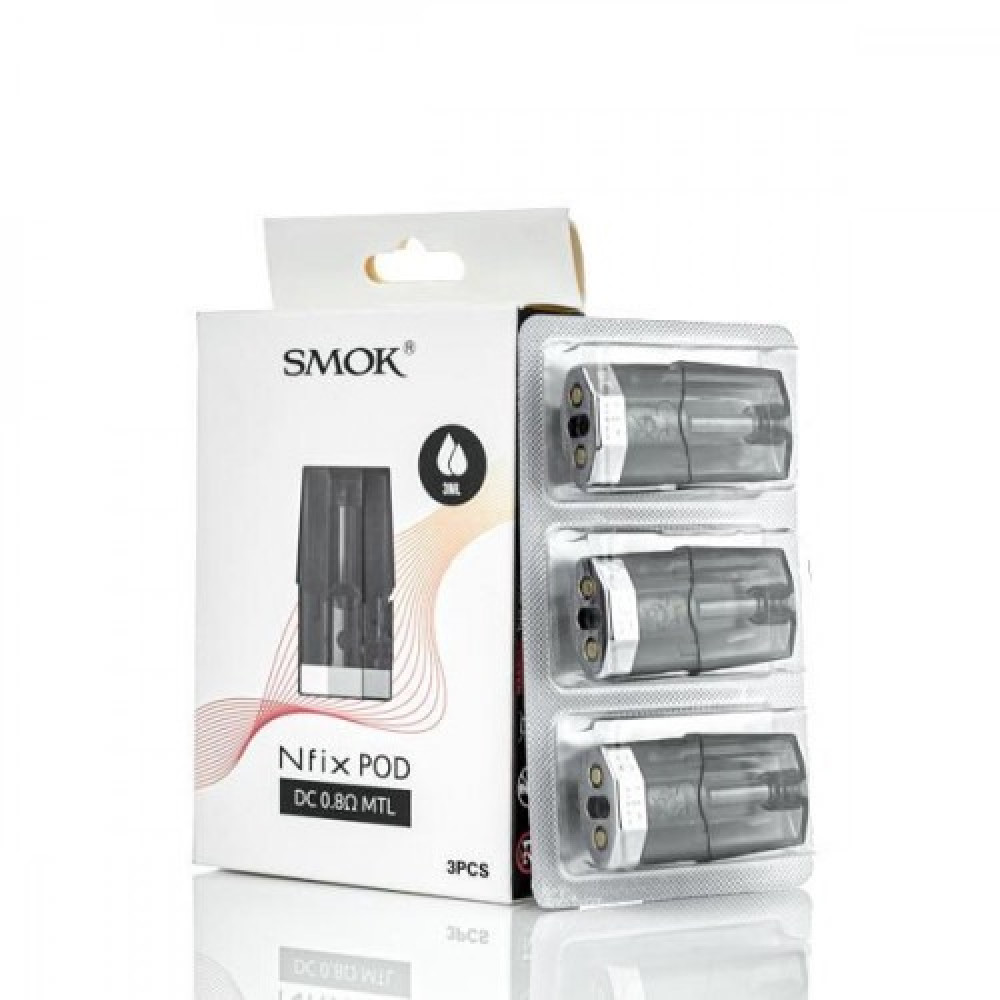
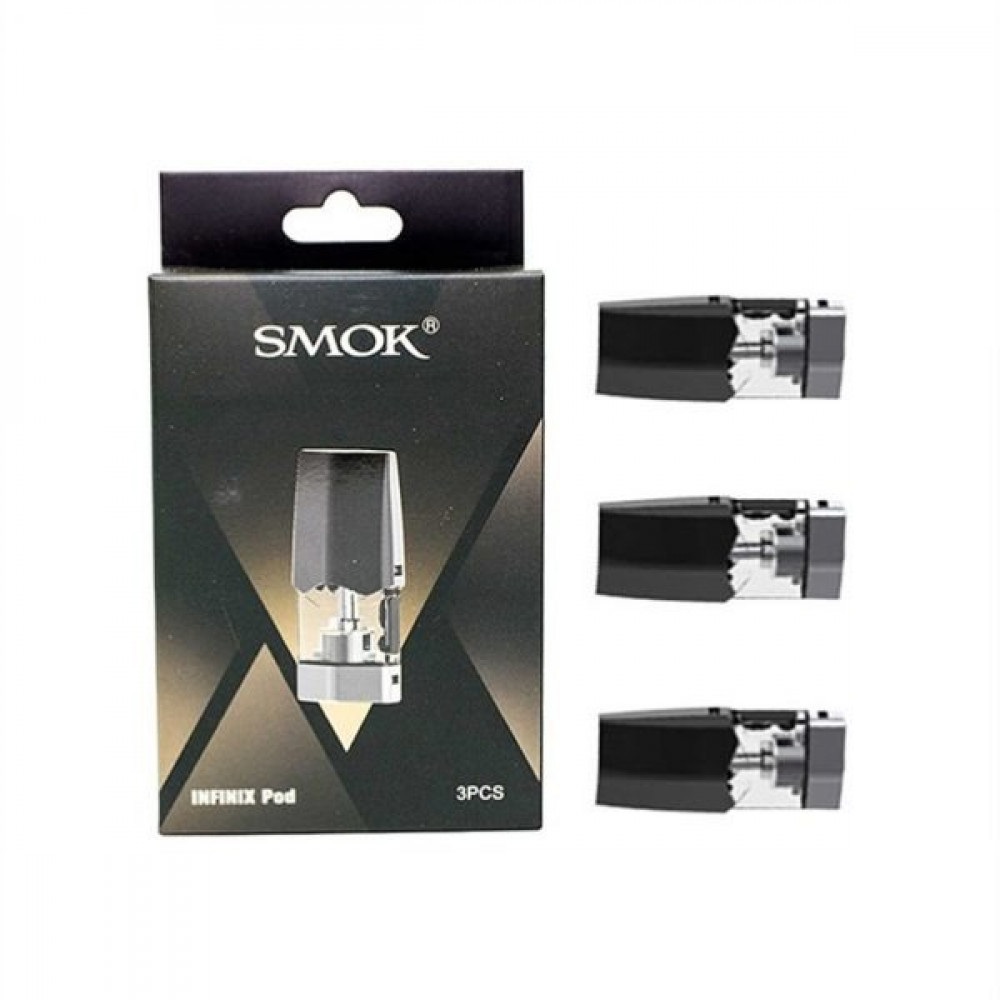
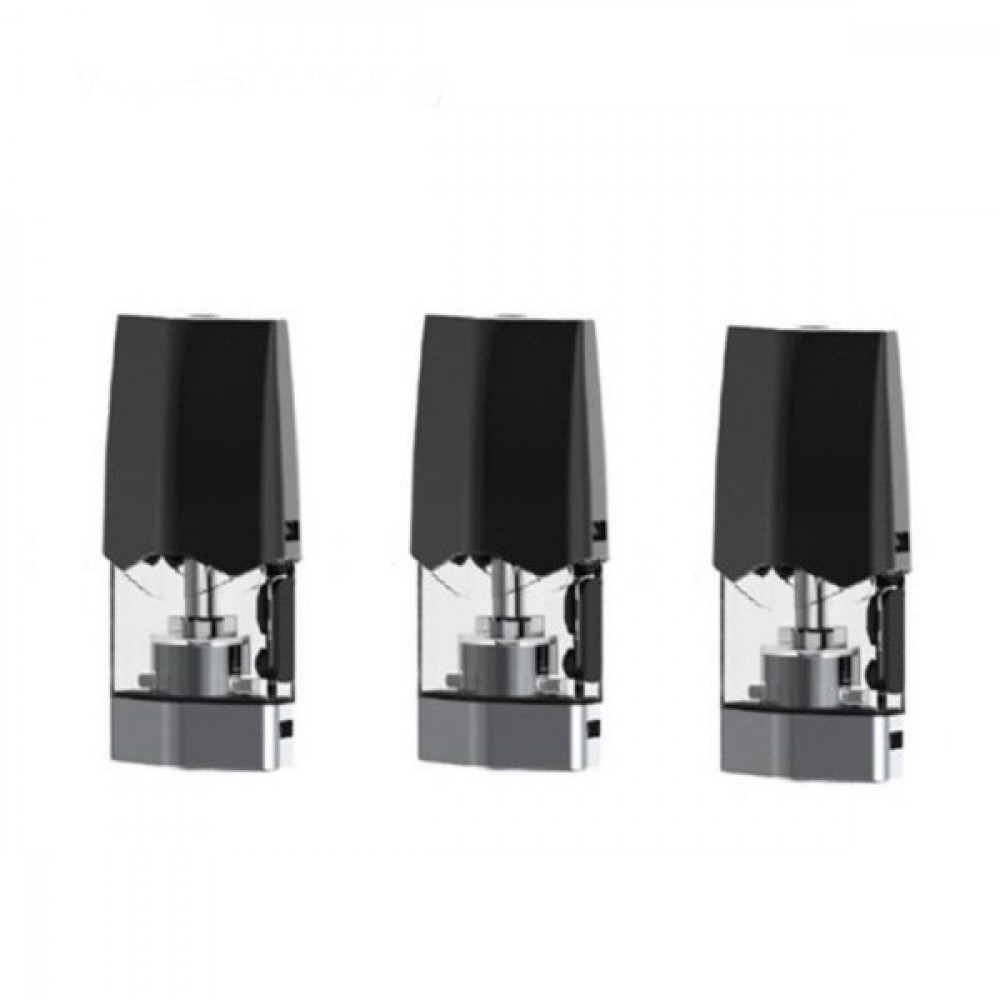
بودات سموك انفنكس الاصدار الاول INFINIX 1 POD - بود بودات POD PODS - فيب كلوب السعودية شيشة الكترونية سحبة سيجارة نكهات
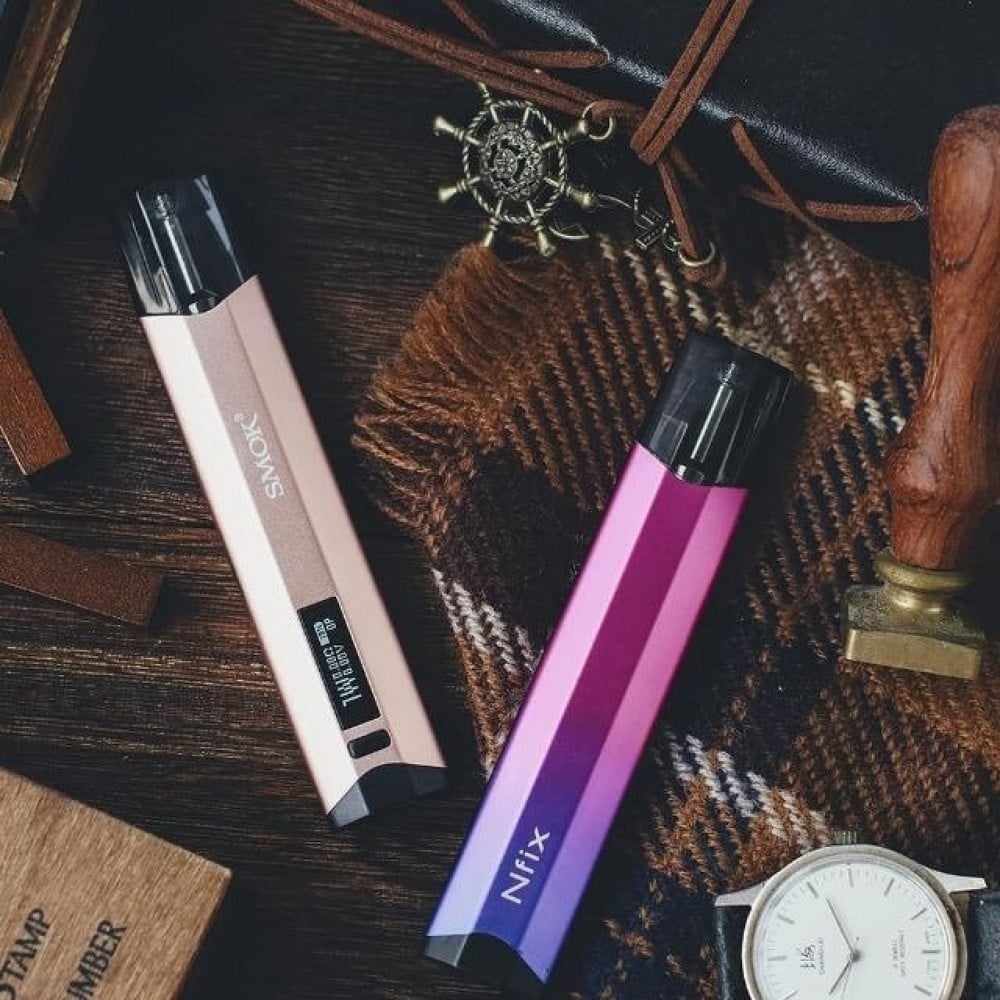

سحبة سموك انفكس كيت Smok NFIX 25W Pod System - متجر زيرو - شيشة الكترونية - - نكهات فيب - فيب السعودية ZERO VAPE

سحبة سموك انفنكس 2 SMOK INFINIX 2 Kit - دخان الكتروني VAPE MYIE SMOK - فيب كلوب السعودية شيشة الكترونية سحبة سيجارة نكهات