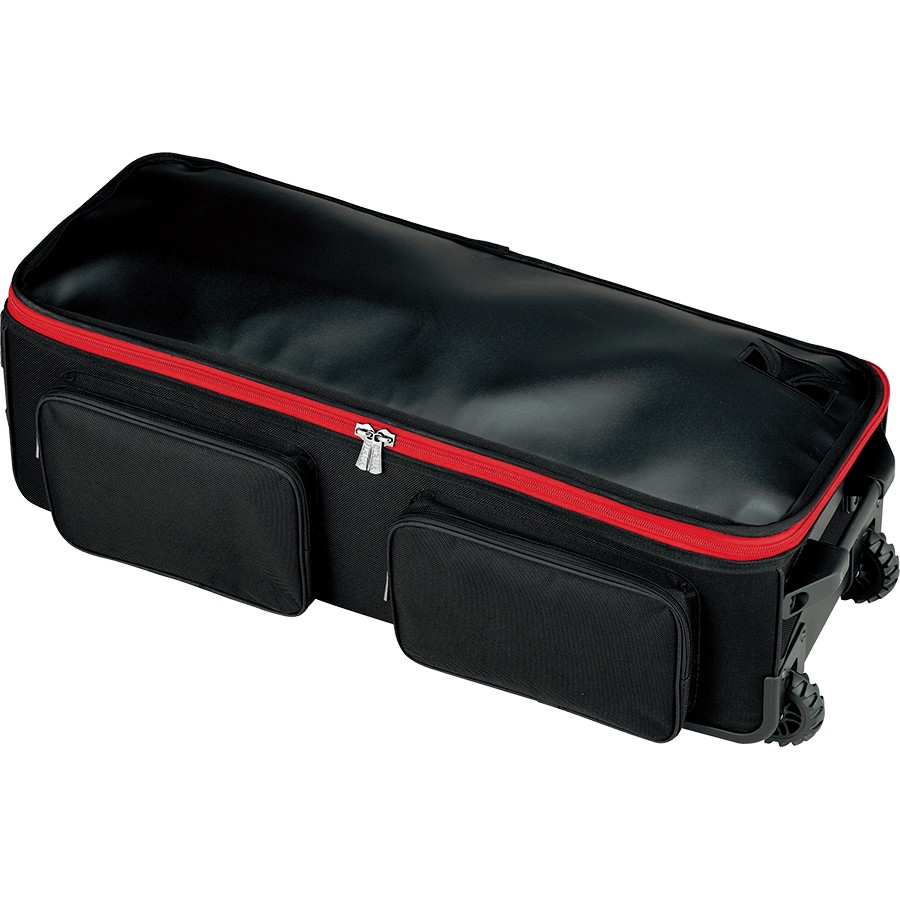
Vásárlás Senrhy Oxford Szövet Acél Nyelv Dob Kézi Táska Tartós Protable Fekete Táska Tárolására Acél Nyelv Dob 10 Es 12 Inch > Kedvezmény < www.lattelife.news

Dob, Doboz, Dob Táska 600d Oxford Szövet, Dob Táska 5mm Tele Pamut Fekete Eszközök: Kahong kedvezmény ~ kedvezmény \ Nagykereskedelem-Eredeti.cam

Kiárusítás! Mini Kerek Női Táska Vad Népszerű Rúzs, Táskák, Divat Textúra Erszényem Gyémánt Lánc Dob Táska Vállon Messenger / Női táskák | ErtekKivezetes.today

Rum Készlet, Kalapács Táska, Marhabőr, Dob Stick Bag, Kényelmes, Szétszerelés, Csirkecomb, Borító Kiárusítás! \ Bolt < Mart-Premium.cam

Rendelés Hordozható eva halászati tekercs táska védőtok dob / spin / tutaj halászati tekercs táska halászati tartozékok ~ Halászati / Timooo.co

Többlet Vietnami Háború Kínai Katonai Dob Magazin Tok Messenger Lőszer Táska - CN022 > eladó \ www.ofeg.info

Motoros Tok Lovaglás Vízálló Nyeregtáska Mochilas Para Moto Tank Táska Hátsó ülésen Táskák Farok Táskák Motoros Csomagtér Dob Táska kedvezmény / Kedvezmény | Elemeket-Gyar.today

Vásárlás Senrhy Oxford Szövet Acél Nyelv Dob Kézi Táska Tartós Protable Fekete Táska Tárolására Acél Nyelv Dob 10 Es 12 Inch > Kedvezmény < www.lattelife.news

Fox Camolite Pót Dob Tartó Táska • Rácvárosi horgászbolt Pécs - horgászbotok, orsók és horgászfelszerelések webáruháza

Disney Aranyos Rajzfilm Mickey Egér Nyomtatott Messenger, Táska, Pénztárca Mickey Válltáska 2020-as Tavaszi Dob Táska Mini Lánc Táska kedvezmény < Babák & Plüss Játékok - Stock-Depot.cam

Fekete vastag oxford szövet, hülye dob táska hordozható pergő botot, bot kotta gyakorlat pad ütősök tartozékok rendelés - Pláza ~ Termekek-Raktar.cam

Kiárusítás Alsócomb Pack Táska Vastag Vízhatlan Oxford Elektronikus Dob Stick Esetben Tud A4-es Eredmények / Outlet - Cheap-Purchase.cyou