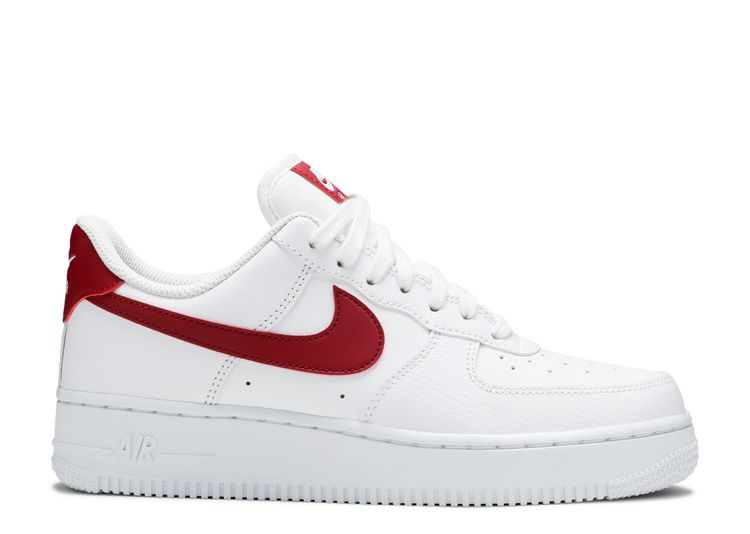
Wmns Air Force 1 '07 'White Noble Red' - Nike - 315115 154 - white/noble red /white/white | Flight Club

154 - Nike Wmns Air Force 1'07 Low White/Noble Red On Sale 315115 - nike air max 1 denim grey paint black color shoes
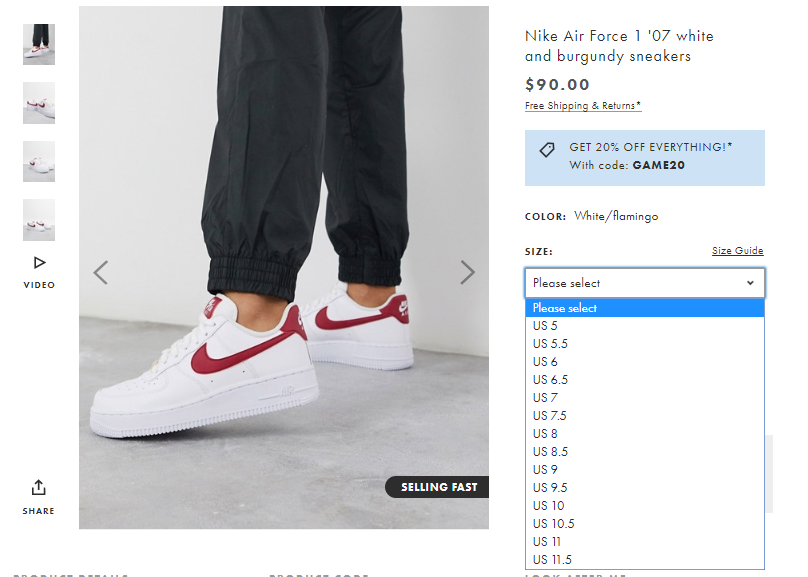
Ballin Sneaks on Twitter: "Nike Air Force 1 Low White/Noble Red 20% off with code GAME20 https://t.co/iEVMAAjnNg https://t.co/Tm0uCcOIKH" / Twitter

Titolo Sneaker Boutique on Instagram: “Nike Wmns Air Force 1 Low in "White/Noble Red". Get Your Pair o… | Nike shoes air force, Air force one shoes, Air force shoes

Buy 2021 Nike Air Force 1 07 White Noble Red Sneakers 315115-154 – Cadysneaker | Sneaker New / Release Info

154 Sneakers – Ietp - nike sb dunk low premium chirping bird sound - Latest Nike nike air retaliate low income women breastfeeding Murky Cringing Sanguine 315115

154 - Nike Wmns Air Force 1'07 Low White/Noble Red On Sale 315115 - nike air penny metallic silver sneakers with jeans

















