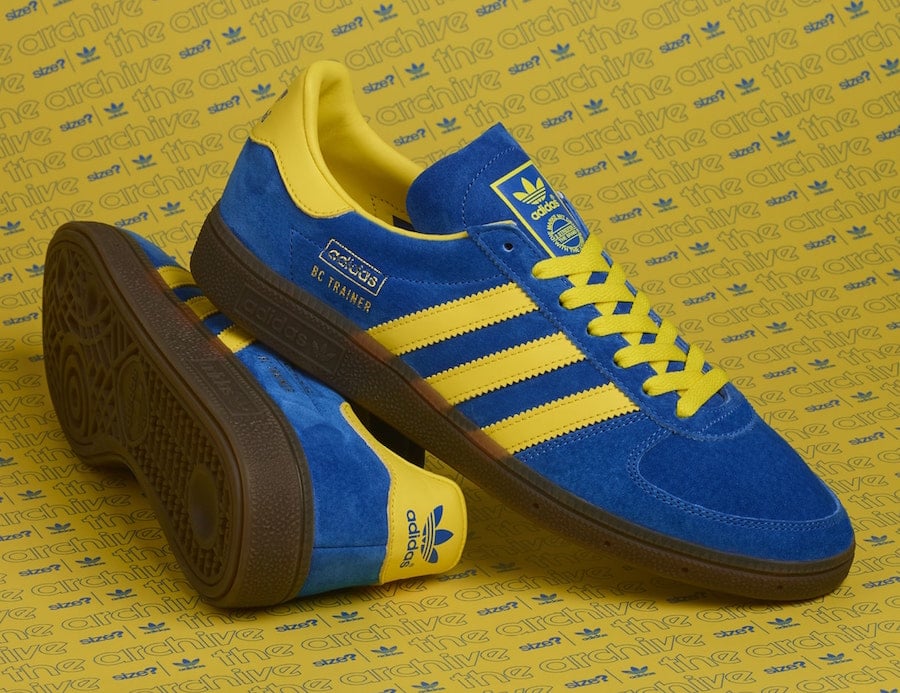
Back From The Archives: adidas Originals Training PT – size? Exclusive | Vintage sneakers, New sneakers, Adidas

adidas Performance STELLA COURT - Tenisice za sve terene - signal green/core black/footwear white/neonski zeleno - Zalando.hr

adidas stadium team backpack soccer tournament - Best shoes JofemarShops - adidas Originals Adicolor Split Trefoil Sweatshirt HC4622