Anvelope Iasi - magazin de anvelope in Iasi • cauciucuri ieftine si oferte pret la anvelope iasi - magazin de anvelope in iasi
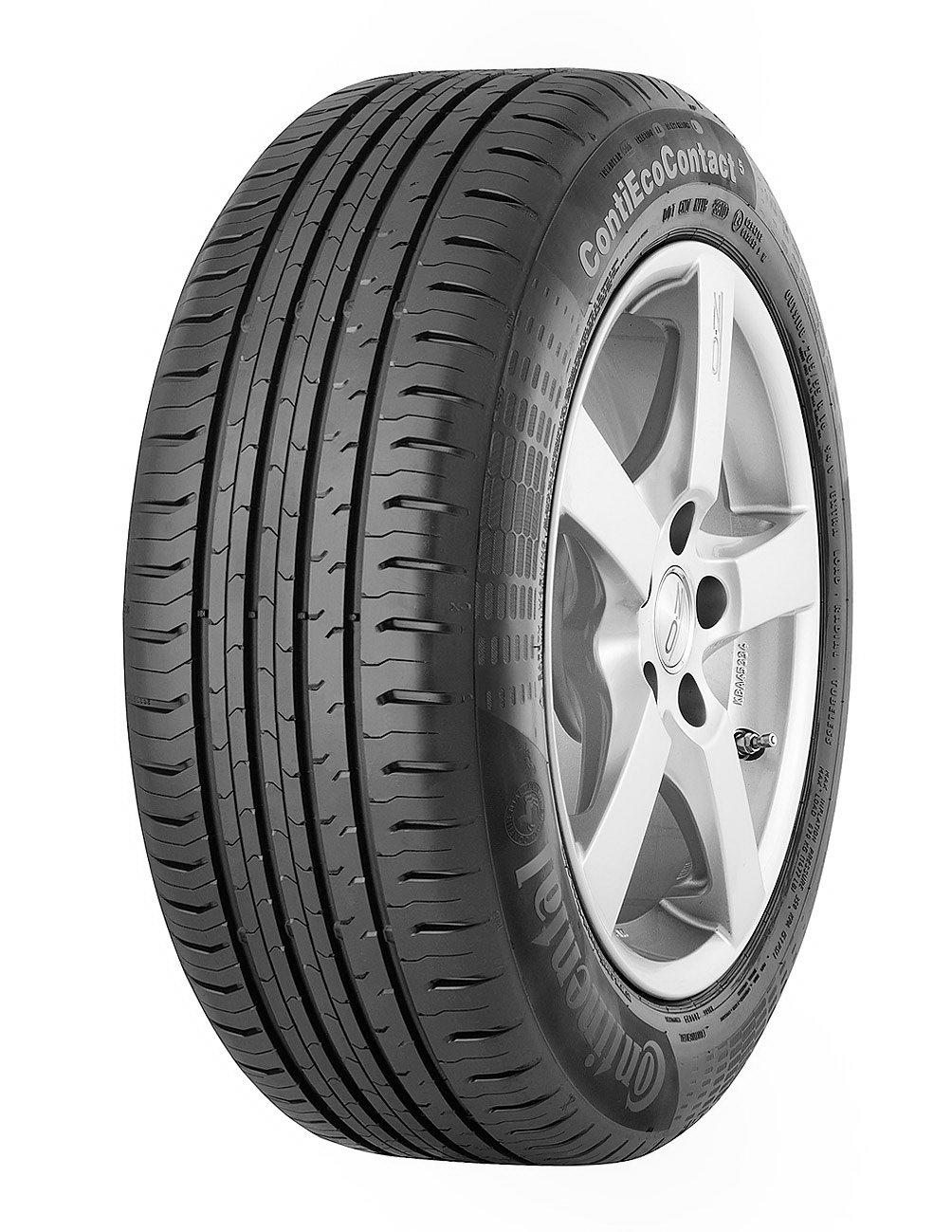

Anvelope vara CONTINENTAL ECO CONTACT 5 205/55 R16 91H • cauciucuri ieftine si oferte pret la anvelope vara continental eco contact 5 205/55 r16 91h